第 13 章 dplyr进阶
本章主要关注dplyr的一些应用。
13.2 变量含义
| variable | class | description |
|---|---|---|
| species | character | 企鹅种类 (Adelie, Gentoo, Chinstrap) |
| island | character | 所在岛屿 (Biscoe, Dream, Torgersen) |
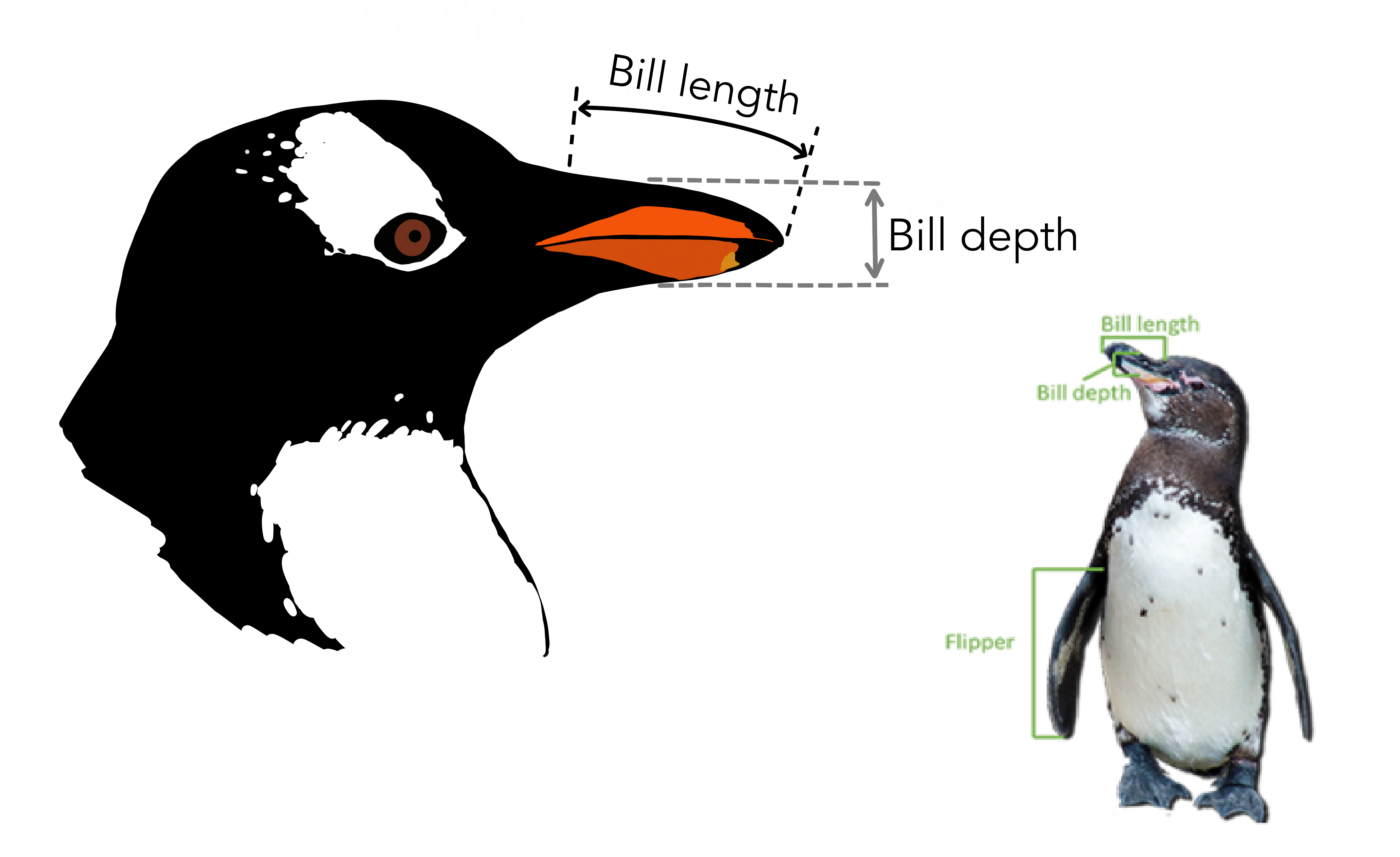
| bill_length_mm | double | 嘴峰长度 (单位毫米) |
| bill_depth_mm | double | 嘴峰深度 (单位毫米) |
| flipper_length_mm | integer | 鰭肢长度 (单位毫米) |
| body_mass_g | integer | 体重 (单位克) |
| sex | character | 性别 |
| year | integer | 记录年份 |
13.3 简单回顾
13.3.1 选择”bill_“开始的列
penguins %>% select(starts_with("bill_"))## # A tibble: 333 × 2
## bill_length_mm bill_depth_mm
## <dbl> <dbl>
## 1 39.1 18.7
## 2 39.5 17.4
## 3 40.3 18
## 4 36.7 19.3
## 5 39.3 20.6
## 6 38.9 17.8
## 7 39.2 19.6
## 8 41.1 17.6
## 9 38.6 21.2
## 10 34.6 21.1
## # ℹ 323 more rows13.3.2 选择”_mm”结尾的列
## # A tibble: 333 × 3
## bill_length_mm bill_depth_mm flipper_length_mm
## <dbl> <dbl> <int>
## 1 39.1 18.7 181
## 2 39.5 17.4 186
## 3 40.3 18 195
## 4 36.7 19.3 193
## 5 39.3 20.6 190
## 6 38.9 17.8 181
## 7 39.2 19.6 195
## 8 41.1 17.6 182
## 9 38.6 21.2 191
## 10 34.6 21.1 198
## # ℹ 323 more rows13.3.3 选择含有”length”的列
## # A tibble: 333 × 2
## bill_length_mm flipper_length_mm
## <dbl> <int>
## 1 39.1 181
## 2 39.5 186
## 3 40.3 195
## 4 36.7 193
## 5 39.3 190
## 6 38.9 181
## 7 39.2 195
## 8 41.1 182
## 9 38.6 191
## 10 34.6 198
## # ℹ 323 more rows13.3.4 选择数值型的列
## # A tibble: 333 × 5
## bill_length_mm bill_depth_mm flipper_length_mm body_mass_g year
## <dbl> <dbl> <int> <int> <int>
## 1 39.1 18.7 181 3750 2007
## 2 39.5 17.4 186 3800 2007
## 3 40.3 18 195 3250 2007
## 4 36.7 19.3 193 3450 2007
## 5 39.3 20.6 190 3650 2007
## 6 38.9 17.8 181 3625 2007
## 7 39.2 19.6 195 4675 2007
## 8 41.1 17.6 182 3200 2007
## 9 38.6 21.2 191 3800 2007
## 10 34.6 21.1 198 4400 2007
## # ℹ 323 more rows13.3.6 选择字符串类型以外的列
## # A tibble: 333 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.5 17.4 186 3800
## 3 Adelie Torgersen 40.3 18 195 3250
## 4 Adelie Torgersen 36.7 19.3 193 3450
## 5 Adelie Torgersen 39.3 20.6 190 3650
## 6 Adelie Torgersen 38.9 17.8 181 3625
## 7 Adelie Torgersen 39.2 19.6 195 4675
## 8 Adelie Torgersen 41.1 17.6 182 3200
## 9 Adelie Torgersen 38.6 21.2 191 3800
## 10 Adelie Torgersen 34.6 21.1 198 4400
## # ℹ 323 more rows
## # ℹ 2 more variables: sex <fct>, year <int>13.3.7 可以用多种组合来选择
penguins %>% select(species, starts_with("bill_"))## # A tibble: 333 × 3
## species bill_length_mm bill_depth_mm
## <fct> <dbl> <dbl>
## 1 Adelie 39.1 18.7
## 2 Adelie 39.5 17.4
## 3 Adelie 40.3 18
## 4 Adelie 36.7 19.3
## 5 Adelie 39.3 20.6
## 6 Adelie 38.9 17.8
## 7 Adelie 39.2 19.6
## 8 Adelie 41.1 17.6
## 9 Adelie 38.6 21.2
## 10 Adelie 34.6 21.1
## # ℹ 323 more rows13.3.9 选择全部为0的列
## # A tibble: 5 × 4
## x y z w
## <dbl> <dbl> <dbl> <dbl>
## 1 1 0 -1 0
## 2 2 0 -2 1
## 3 3 0 0 2
## 4 4 0 2 3
## 5 5 0 1 4## # A tibble: 5 × 1
## y
## <dbl>
## 1 0
## 2 0
## 3 0
## 4 0
## 5 0## # A tibble: 5 × 1
## y
## <dbl>
## 1 0
## 2 0
## 3 0
## 4 0
## 5 0## # A tibble: 5 × 2
## y z
## <dbl> <dbl>
## 1 0 -1
## 2 0 -2
## 3 0 0
## 4 0 2
## 5 0 1## # A tibble: 5 × 3
## y z w
## <dbl> <dbl> <dbl>
## 1 0 -1 0
## 2 0 -2 1
## 3 0 0 2
## 4 0 2 3
## 5 0 1 4课堂练习:剔除全部为NA的列或者全部为NA的行
df <- tibble(
x = c(NA, NA, NA),
y = c(2, 3, NA),
z = c(NA, 5, NA)
)
# columns
df %>%
select(where(~ !all(is.na(.x))))## # A tibble: 3 × 2
## y z
## <dbl> <dbl>
## 1 2 NA
## 2 3 5
## 3 NA NA
# rows
df %>%
filter(
if_any(everything(), ~ !is.na(.x))
)## # A tibble: 2 × 3
## x y z
## <lgl> <dbl> <dbl>
## 1 NA 2 NA
## 2 NA 3 513.3.10 寻找男企鹅
函数 filter() 中的逻辑运算符
| Operator | Meaning |
|---|---|
== |
Equal to |
> |
Greater than |
< |
Less than |
>= |
Greater than or equal to |
<= |
Less than or equal to |
!= |
Not equal to |
%in% |
in |
is.na |
is a missing value (NA) |
!is.na |
is not a missing value |
& |
and |
| |
or |
## # A tibble: 168 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.3 20.6 190 3650
## 3 Adelie Torgersen 39.2 19.6 195 4675
## 4 Adelie Torgersen 38.6 21.2 191 3800
## 5 Adelie Torgersen 34.6 21.1 198 4400
## 6 Adelie Torgersen 42.5 20.7 197 4500
## 7 Adelie Torgersen 46 21.5 194 4200
## 8 Adelie Biscoe 37.7 18.7 180 3600
## 9 Adelie Biscoe 38.2 18.1 185 3950
## 10 Adelie Biscoe 38.8 17.2 180 3800
## # ℹ 158 more rows
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 265 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.5 17.4 186 3800
## 3 Adelie Torgersen 40.3 18 195 3250
## 4 Adelie Torgersen 36.7 19.3 193 3450
## 5 Adelie Torgersen 39.3 20.6 190 3650
## 6 Adelie Torgersen 38.9 17.8 181 3625
## 7 Adelie Torgersen 39.2 19.6 195 4675
## 8 Adelie Torgersen 41.1 17.6 182 3200
## 9 Adelie Torgersen 38.6 21.2 191 3800
## 10 Adelie Torgersen 34.6 21.1 198 4400
## # ℹ 255 more rows
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 50 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 40.3 18 195 3250
## 2 Adelie Torgersen 41.1 17.6 182 3200
## 3 Adelie Torgersen 42.5 20.7 197 4500
## 4 Adelie Torgersen 46 21.5 194 4200
## 5 Adelie Biscoe 40.6 18.6 183 3550
## 6 Adelie Biscoe 40.5 17.9 187 3200
## 7 Adelie Biscoe 40.5 18.9 180 3950
## 8 Adelie Dream 40.9 18.9 184 3900
## 9 Adelie Dream 42.2 18.5 180 3550
## 10 Adelie Dream 40.8 18.4 195 3900
## # ℹ 40 more rows
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 50 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 40.3 18 195 3250
## 2 Adelie Torgersen 41.1 17.6 182 3200
## 3 Adelie Torgersen 42.5 20.7 197 4500
## 4 Adelie Torgersen 46 21.5 194 4200
## 5 Adelie Biscoe 40.6 18.6 183 3550
## 6 Adelie Biscoe 40.5 17.9 187 3200
## 7 Adelie Biscoe 40.5 18.9 180 3950
## 8 Adelie Dream 40.9 18.9 184 3900
## 9 Adelie Dream 42.2 18.5 180 3550
## 10 Adelie Dream 40.8 18.4 195 3900
## # ℹ 40 more rows
## # ℹ 2 more variables: sex <fct>, year <int>课堂练习,说出以下代码的含义
13.4 更多应用
希望介绍一个技术,对应一个应用场景
13.4.1 弱水三千,只取一瓢
## # A tibble: 6 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.5 17.4 186 3800
## 3 Adelie Torgersen 40.3 18 195 3250
## 4 Adelie Torgersen 36.7 19.3 193 3450
## 5 Adelie Torgersen 39.3 20.6 190 3650
## 6 Adelie Torgersen 38.9 17.8 181 3625
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 6 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Chinstrap Dream 45.7 17 195 3650
## 2 Chinstrap Dream 55.8 19.8 207 4000
## 3 Chinstrap Dream 43.5 18.1 202 3400
## 4 Chinstrap Dream 49.6 18.2 193 3775
## 5 Chinstrap Dream 50.8 19 210 4100
## 6 Chinstrap Dream 50.2 18.7 198 3775
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 1 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 3 × 8
## # Groups: species [3]
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Chinstrap Dream 46.5 17.9 192 3500
## 3 Gentoo Biscoe 46.1 13.2 211 4500
## # ℹ 2 more variables: sex <fct>, year <int>13.4.2 嘴峰长度最大那一行
三种方法
## # A tibble: 1 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Gentoo Biscoe 59.6 17 230 6050
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 1 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Gentoo Biscoe 59.6 17 230 6050
## # ℹ 2 more variables: sex <fct>, year <int>## # A tibble: 1 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Gentoo Biscoe 59.6 17 230 6050
## # ℹ 2 more variables: sex <fct>, year <int>13.4.3 separate
tb <- tibble::tribble(
~day, ~price,
1, "30-45",
2, "40-95",
3, "89-65",
4, "45-63",
5, "52-42"
)## # A tibble: 5 × 3
## day low high
## <dbl> <chr> <chr>
## 1 1 30 45
## 2 2 40 95
## 3 3 89 65
## 4 4 45 63
## 5 5 52 4213.4.4 unite
## # A tibble: 5 × 4
## day price low high
## <dbl> <chr> <chr> <chr>
## 1 1 30:45 30 45
## 2 2 40:95 40 95
## 3 3 89:65 89 65
## 4 4 45:63 45 63
## 5 5 52:42 52 4213.4.5 distinct
distinct()处理的对象是data.frame;功能是筛选不重复的row;返回data.frame
df <- tibble::tribble(
~x, ~y, ~z,
1, 1, 1,
1, 1, 2,
1, 1, 1,
2, 1, 2,
2, 2, 3,
3, 3, 1
)
df## # A tibble: 6 × 3
## x y z
## <dbl> <dbl> <dbl>
## 1 1 1 1
## 2 1 1 2
## 3 1 1 1
## 4 2 1 2
## 5 2 2 3
## 6 3 3 1## # A tibble: 5 × 3
## x y z
## <dbl> <dbl> <dbl>
## 1 1 1 1
## 2 1 1 2
## 3 2 1 2
## 4 2 2 3
## 5 3 3 1## # A tibble: 3 × 1
## x
## <dbl>
## 1 1
## 2 2
## 3 3## # A tibble: 4 × 2
## x y
## <dbl> <dbl>
## 1 1 1
## 2 2 1
## 3 2 2
## 4 3 3## # A tibble: 4 × 3
## x y z
## <dbl> <dbl> <dbl>
## 1 1 1 1
## 2 2 1 2
## 3 2 2 3
## 4 3 3 1## # A tibble: 4 × 3
## # Groups: x [3]
## x y z
## <dbl> <dbl> <dbl>
## 1 1 1 1
## 2 2 1 2
## 3 2 2 3
## 4 3 3 1n_distinct()处理的对象是vector;功能是统计不同的元素有多少个;返回一个数值
c(1, 1, 1, 2, 2, 1, 3, 3) %>% n_distinct()## [1] 3
df$z %>% n_distinct()## [1] 3
df %>%
group_by(x) %>%
summarise(
n = n_distinct(z)
)## # A tibble: 3 × 2
## x n
## <dbl> <int>
## 1 1 2
## 2 2 2
## 3 3 113.4.6 有关NA的计算
NA很讨厌,凡是它参与的四则运算,结果都是NA,
## [1] NA所以需要事先把它删除,增加参数说明 na.rm = TRUE
## [1] 7## [1] 2.33333313.4.7 寻找企鹅中的胖子
## # A tibble: 333 × 9
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.5 17.4 186 3800
## 3 Adelie Torgersen 40.3 18 195 3250
## 4 Adelie Torgersen 36.7 19.3 193 3450
## 5 Adelie Torgersen 39.3 20.6 190 3650
## 6 Adelie Torgersen 38.9 17.8 181 3625
## 7 Adelie Torgersen 39.2 19.6 195 4675
## 8 Adelie Torgersen 41.1 17.6 182 3200
## 9 Adelie Torgersen 38.6 21.2 191 3800
## 10 Adelie Torgersen 34.6 21.1 198 4400
## # ℹ 323 more rows
## # ℹ 3 more variables: sex <fct>, year <int>, body <chr>随堂练习:用考试成绩的均值代替缺失值
df <- tibble::tribble(
~name, ~type, ~score,
"Alice", "english", 80,
"Alice", "math", NA,
"Bob", "english", 70,
"Bob", "math", 69,
"Carol", "english", NA,
"Carol", "math", 90
)
df## # A tibble: 6 × 3
## name type score
## <chr> <chr> <dbl>
## 1 Alice english 80
## 2 Alice math NA
## 3 Bob english 70
## 4 Bob math 69
## 5 Carol english NA
## 6 Carol math 90
df %>%
group_by(type) %>%
mutate(mean_score = mean(score, na.rm = TRUE)) %>%
mutate(newscore = if_else(is.na(score), mean_score, score))## # A tibble: 6 × 5
## # Groups: type [2]
## name type score mean_score newscore
## <chr> <chr> <dbl> <dbl> <dbl>
## 1 Alice english 80 75 80
## 2 Alice math NA 79.5 79.5
## 3 Bob english 70 75 70
## 4 Bob math 69 79.5 69
## 5 Carol english NA 75 75
## 6 Carol math 90 79.5 9013.4.8 给企鹅身材分类
penguins %>%
mutate(
body = case_when(
body_mass_g < 3500 ~ "best",
body_mass_g >= 3500 & body_mass_g < 4500 ~ "good",
body_mass_g >= 4500 & body_mass_g < 5500 ~ "general",
TRUE ~ "other"
)
)## # A tibble: 333 × 9
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.5 17.4 186 3800
## 3 Adelie Torgersen 40.3 18 195 3250
## 4 Adelie Torgersen 36.7 19.3 193 3450
## 5 Adelie Torgersen 39.3 20.6 190 3650
## 6 Adelie Torgersen 38.9 17.8 181 3625
## 7 Adelie Torgersen 39.2 19.6 195 4675
## 8 Adelie Torgersen 41.1 17.6 182 3200
## 9 Adelie Torgersen 38.6 21.2 191 3800
## 10 Adelie Torgersen 34.6 21.1 198 4400
## # ℹ 323 more rows
## # ℹ 3 more variables: sex <fct>, year <int>, body <chr>随堂练习:按嘴峰长度分成A, B, C, D 4个等级
penguins %>%
mutate(
degree = case_when(
bill_length_mm < 35 ~ "A",
bill_length_mm >= 35 & bill_length_mm < 45 ~ "B",
bill_length_mm >= 45 & bill_length_mm < 55 ~ "C",
TRUE ~ "D"
)
)## # A tibble: 333 × 9
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.5 17.4 186 3800
## 3 Adelie Torgersen 40.3 18 195 3250
## 4 Adelie Torgersen 36.7 19.3 193 3450
## 5 Adelie Torgersen 39.3 20.6 190 3650
## 6 Adelie Torgersen 38.9 17.8 181 3625
## 7 Adelie Torgersen 39.2 19.6 195 4675
## 8 Adelie Torgersen 41.1 17.6 182 3200
## 9 Adelie Torgersen 38.6 21.2 191 3800
## 10 Adelie Torgersen 34.6 21.1 198 4400
## # ℹ 323 more rows
## # ℹ 3 more variables: sex <fct>, year <int>, degree <chr>13.4.9 每种类型企鹅有多少只?
知识点:n()函数,统计当前分组数据框的行数
## # A tibble: 1 × 1
## n
## <int>
## 1 333## # A tibble: 3 × 2
## species n
## <fct> <int>
## 1 Adelie 146
## 2 Chinstrap 68
## 3 Gentoo 119统计某个变量中各组出现的次数,可以使用count()函数
## # A tibble: 3 × 2
## species n
## <fct> <int>
## 1 Adelie 146
## 2 Chinstrap 68
## 3 Gentoo 119不同性别的企鹅各有多少
## # A tibble: 2 × 2
## sex n
## <fct> <int>
## 1 male 168
## 2 female 165可以统计不同组合出现的次数
## # A tibble: 5 × 3
## island species n
## <fct> <fct> <int>
## 1 Biscoe Adelie 44
## 2 Biscoe Gentoo 119
## 3 Dream Adelie 55
## 4 Dream Chinstrap 68
## 5 Torgersen Adelie 47可以在count()里构建新变量,并利用这个新变量完成统计。
比如,统计嘴巴长度大于40的企鹅个数
- 常规做法
## # A tibble: 1 × 1
## n
## <int>
## 1 237-
count()做法
## # A tibble: 2 × 2
## longer_bill n
## <lgl> <int>
## 1 FALSE 96
## 2 TRUE 237解析思路:bill_length_mm > 40 比较算符 返回逻辑型向量,向量里面只有TRUR和FALSE两种值,因此上面的代码相当于统计TRUE有多少个,FALSE有多少个?
13.4.10 强制转换
矢量中的元素必须是相同的类型,但如果不一样呢,会发生什么? 这个时候R会强制转换成相同的类型。这就涉及数据类型的转换层级
- character > numeric > logical
- double > integer
比如这里会强制转换成字符串类型
c("foo", 1, TRUE)## [1] "foo" "1" "TRUE"这里会强制转换成数值型
c(1, TRUE, FALSE)## [1] 1 1 0## [1] 2随堂练习:补全下面代码,求嘴峰长度大于40mm的占比?
## # A tibble: 333 × 9
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen 39.1 18.7 181 3750
## 2 Adelie Torgersen 39.5 17.4 186 3800
## 3 Adelie Torgersen 40.3 18 195 3250
## 4 Adelie Torgersen 36.7 19.3 193 3450
## 5 Adelie Torgersen 39.3 20.6 190 3650
## 6 Adelie Torgersen 38.9 17.8 181 3625
## 7 Adelie Torgersen 39.2 19.6 195 4675
## 8 Adelie Torgersen 41.1 17.6 182 3200
## 9 Adelie Torgersen 38.6 21.2 191 3800
## 10 Adelie Torgersen 34.6 21.1 198 4400
## # ℹ 323 more rows
## # ℹ 3 more variables: sex <fct>, year <int>, is_bigger40 <lgl>13.5 across()之美
我们想知道,嘴巴长度和厚度的均值
## # A tibble: 1 × 1
## length
## <dbl>
## 1 44.0接着添加下个变量
## # A tibble: 1 × 2
## length depth
## <dbl> <dbl>
## 1 44.0 44.0长度和厚度惊人的相等。我是不是发现新大陆了?
13.5.1 across()函数
更安全、更简练的写法,王老师的最爱
## # A tibble: 1 × 2
## bill_depth_mm bill_length_mm
## <dbl> <dbl>
## 1 17.2 44.0翅膀的长度加进去看看
## # A tibble: 1 × 3
## bill_depth_mm bill_length_mm flipper_length_mm
## <dbl> <dbl> <dbl>
## 1 17.2 44.0 201.还可以更简练喔
## # A tibble: 1 × 3
## bill_length_mm bill_depth_mm flipper_length_mm
## <dbl> <dbl> <dbl>
## 1 44.0 17.2 201.across()函数用法
across(.cols = everything(), .fns = NULL, ..., .names = NULL)- 用在
mutate()和summarise()函数里面 -
across()对多列执行相同的函数操作,返回数据框
13.5.2 数据中心化
penguins %>%
mutate(
bill_length_mm = bill_length_mm - mean(bill_length_mm),
bill_depth_mm = bill_depth_mm - mean(bill_depth_mm)
)## # A tibble: 333 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen -4.89 1.54 181 3750
## 2 Adelie Torgersen -4.49 0.235 186 3800
## 3 Adelie Torgersen -3.69 0.835 195 3250
## 4 Adelie Torgersen -7.29 2.14 193 3450
## 5 Adelie Torgersen -4.69 3.44 190 3650
## 6 Adelie Torgersen -5.09 0.635 181 3625
## 7 Adelie Torgersen -4.79 2.44 195 4675
## 8 Adelie Torgersen -2.89 0.435 182 3200
## 9 Adelie Torgersen -5.39 4.04 191 3800
## 10 Adelie Torgersen -9.39 3.94 198 4400
## # ℹ 323 more rows
## # ℹ 2 more variables: sex <fct>, year <int>更清晰的办法
centralized <- function(x) {
x - mean(x)
}
penguins %>%
mutate(
across(c(bill_length_mm, bill_depth_mm), centralized)
)## # A tibble: 333 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen -4.89 1.54 181 3750
## 2 Adelie Torgersen -4.49 0.235 186 3800
## 3 Adelie Torgersen -3.69 0.835 195 3250
## 4 Adelie Torgersen -7.29 2.14 193 3450
## 5 Adelie Torgersen -4.69 3.44 190 3650
## 6 Adelie Torgersen -5.09 0.635 181 3625
## 7 Adelie Torgersen -4.79 2.44 195 4675
## 8 Adelie Torgersen -2.89 0.435 182 3200
## 9 Adelie Torgersen -5.39 4.04 191 3800
## 10 Adelie Torgersen -9.39 3.94 198 4400
## # ℹ 323 more rows
## # ℹ 2 more variables: sex <fct>, year <int>13.5.3 数据标准化
std <- function(x) {
(x - mean(x, na.rm = TRUE)) / sd(x, na.rm = TRUE)
}
penguins %>%
mutate(
across(c(bill_length_mm, bill_depth_mm), std)
)## # A tibble: 333 × 8
## species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
## <fct> <fct> <dbl> <dbl> <int> <int>
## 1 Adelie Torgersen -0.895 0.780 181 3750
## 2 Adelie Torgersen -0.822 0.119 186 3800
## 3 Adelie Torgersen -0.675 0.424 195 3250
## 4 Adelie Torgersen -1.33 1.08 193 3450
## 5 Adelie Torgersen -0.858 1.74 190 3650
## 6 Adelie Torgersen -0.931 0.323 181 3625
## 7 Adelie Torgersen -0.876 1.24 195 4675
## 8 Adelie Torgersen -0.529 0.221 182 3200
## 9 Adelie Torgersen -0.986 2.05 191 3800
## 10 Adelie Torgersen -1.72 2.00 198 4400
## # ℹ 323 more rows
## # ℹ 2 more variables: sex <fct>, year <int>或者使用更简洁的方法
# using across() and purrr style
penguins %>%
summarise(
across(starts_with("bill_"), ~ (.x - mean(.x)) / sd(.x))
)## # A tibble: 333 × 2
## bill_length_mm bill_depth_mm
## <dbl> <dbl>
## 1 -0.895 0.780
## 2 -0.822 0.119
## 3 -0.675 0.424
## 4 -1.33 1.08
## 5 -0.858 1.74
## 6 -0.931 0.323
## 7 -0.876 1.24
## 8 -0.529 0.221
## 9 -0.986 2.05
## 10 -1.72 2.00
## # ℹ 323 more rows13.5.4 多列多个统计函数
penguins %>%
group_by(species) %>%
summarise(
across(ends_with("_mm"), list(mean = mean, sd = sd), na.rm = TRUE)
)## # A tibble: 3 × 7
## species bill_length_mm_mean bill_length_mm_sd bill_depth_mm_mean
## <fct> <dbl> <dbl> <dbl>
## 1 Adelie 38.8 2.66 18.3
## 2 Chinstrap 48.8 3.34 18.4
## 3 Gentoo 47.6 3.11 15.0
## # ℹ 3 more variables: bill_depth_mm_sd <dbl>, flipper_length_mm_mean <dbl>,
## # flipper_length_mm_sd <dbl>随堂练习:以sex分组,对”bill_“开头的列,求出每列的最大值和最小值
penguins %>%
group_by(sex) %>%
summarise(
across(starts_with("bill_"), list(max = max, min = min), na.rm = TRUE)
)## # A tibble: 2 × 5
## sex bill_length_mm_max bill_length_mm_min bill_depth_mm_max
## <fct> <dbl> <dbl> <dbl>
## 1 female 58 32.1 20.7
## 2 male 59.6 34.6 21.5
## # ℹ 1 more variable: bill_depth_mm_min <dbl>