36 Endogeneity
Refresher
A general model framework
Y=Xβ+ϵ
where
Y=n×1
X=n×k
β=k×1
ϵ=n×1
Then, OLS estimates of coefficients are
ˆβOLS=(X′X)−1(X′Y)=(X′X)−1(X′(Xβ+ϵ))=(X′X)−1(X′X)β+(X′X)−1(X′ϵ)ˆβOLS→β+(X′X)−1(X′ϵ)
To have unbiased estimates, we have to get rid of the second part (X′X)−1(X′ϵ)
There are 2 conditions to achieve unbiased estimates:
- E(ϵ|X)=0 (This is easy, putting an intercept can solve this issue)
- Cov(X,ϵ)=0 (This is the hard part)
We only care about omitted variable
Usually, the problem will stem Omitted Variables Bias, but we only care about omitted variable bias when
- Omitted variables correlate with the variables we care about (X). If OMV does not correlate with X, we don’t care, and random assignment makes this correlation goes to 0)
- Omitted variables correlates with outcome/ dependent variable
There are more types of endogeneity listed below.
Types of endogeneity (See Hill et al. (2021) for a review in management):
-
Omitted Variables Bias
- Motivation
- Ability/talent
- Self-selection
Feedback Effect (Simultaneity): also known as bidirectionality
Reverse Causality: Subtle difference from Simultaneity: Technically, two variables affect each other sequentially, but in a big enough time frame, (e.g., monthly, or yearly), our coefficient will be biased just like simultaneity.
To deal with this problem, we have a toolbox (that has been mentioned in previous chapter ??)
Using control variables in regression is a “selection on observables” identification strategy.
In other words, if you believe you have an omitted variable, and you can measure it, including it in the regression model solves your problem. These uninterested variables are called control variables in your model.
However, this is rarely the case (because the problem is we don’t have their measurements). Hence, we need more elaborate methods:
Before we get to methods that deal with bias arises from omitted variables, we consider cases where we do have measurements of a variable, but there is measurement error (bias).
36.1 Endogenous Treatment
36.1.1 Measurement Error
-
Data error can stem from
Coding errors
Reporting errors
Two forms of measurement error:
- Random (stochastic) (indeterminate error) (Classical Measurement Errors): noise or measurement errors do not show up in a consistent or predictable way.
- Systematic (determinate error) (Non-classical Measurement Errors): When measurement error is consistent and predictable across observations.
- Instrument errors (e.g., faulty scale) -> calibration or adjustment
- Method errors (e.g., sampling errors) -> better method development + study design
- Human errors (e.g., judgement)
Usually the systematic measurement error is a bigger issue because it introduces “bias” into our estimates, while random error introduces noise into our estimates
- Noise -> regression estimate to 0
- Bias -> can pull estimate to upward or downward.
36.1.1.1 Classical Measurement Errors
36.1.1.1.1 Right-hand side
- Right-hand side measurement error: When the measurement is in the covariates, then we have the endogeneity problem.
Say you know the true model is
Yi=β0+β1Xi+ui
But you don’t observe Xi, but you observe
˜Xi=Xi+ei
which is known as classical measurement errors where we assume ei is uncorrelated with Xi (i.e., E(Xiei)=0)
Then, when you estimate your observed variables, you have (substitute Xi with ˜Xi−ei ):
Yi=β0+β1(˜Xi−ei)+ui=β0+β1˜Xi+ui−β1ei=β0+β1˜Xi+vi
In words, the measurement error in Xi is now a part of the error term in the regression equation vi. Hence, we have an endogeneity bias.
Endogeneity arises when
E(˜Xivi)=E((Xi+ei)(ui−β1ei))=−β1Var(ei)≠0
Since ˜Xi and ei are positively correlated, then it leads to
a negative bias in ˆβ1 if the true β1 is positive
a positive bias if β1 is negative
In other words, measurement errors cause attenuation bias, which inter turn pushes the coefficient towards 0
As Var(ei) increases or Var(ei)Var(˜X)→1 then ei is a random (noise) and β1→0 (random variable ˜X should not have any relation to Yi)
Technical note:
The size of the bias in the OLS-estimator is
ˆβOLS=cov(˜X,Y)var(˜X)=cov(X+e,βX+u)var(X+e)
then
plimˆβOLS=βσ2Xσ2X+σ2e=βλ
where λ is reliability or signal-to-total variance ratio or attenuation factor
Reliability affect the extent to which measurement error attenuates ˆβ. The attenuation bias is
ˆβOLS−β=−(1−λ)β
Thus, ˆβOLS<β (unless λ=1, in which case we don’t even have measurement error).
Note:
Data transformation worsen (magnify) the measurement error
y=βx+γx2+ϵ
then, the attenuation factor for ˆγ is the square of the attenuation factor for ˆβ (i.e., λˆγ=λ2ˆβ)
Adding covariates increases attenuation bias
To fix classical measurement error problem, we can
- Find estimates of either σ2X,σ2ϵ or λ from validation studies, or survey data.
- Endogenous Treatment Use instrument Z correlated with X but uncorrelated with ϵ
- Abandon your project
36.1.1.1.2 Left-hand side
When the measurement is in the outcome variable, econometricians or causal scientists do not care because they still have an unbiased estimate of the coefficients (the zero conditional mean assumption is not violated, hence we don’t have endogeneity). However, statisticians might care because it might inflate our uncertainty in the coefficient estimates (i.e., higher standard errors).
˜Y=Y+v
then the model you estimate is
˜Y=βX+u+v
Since v is uncorrelated with X, then ˆβ is consistently estimated by OLS
If we have measurement error in Yi, it will pass through β1 and go to ui
36.1.1.2 Non-classical Measurement Errors
Relaxing the assumption that X and ϵ are uncorrelated
Recall the true model we have true estimate is
ˆβ=cov(X+ϵ,βX+u)var(X+ϵ)
then without the above assumption, we have
plimˆβ=β(σ2X+σXϵ)σ2X+σ2ϵ+2σXϵ=(1−σ2ϵ+σXϵσ2X+σ2ϵ+2σXϵ)β=(1−bϵ˜X)β
where bϵ˜X is the covariance between ˜X and ϵ (also the regression coefficient of a regression of ϵ on ˜X)
Hence, the Classical Measurement Errors is just a special case of Non-classical Measurement Errors where bϵ˜X=1−λ
So when σXϵ=0 (Classical Measurement Errors), increasing this covariance bϵ˜X increases the covariance increases the attenuation factor if more than half of the variance in ˜X is measurement error, and decreases the attenuation factor otherwise. This is also known as mean reverting measurement error Bound, Brown, and Mathiowetz (2001)
A general framework for both right-hand side and left-hand side measurement error is (Bound, Brown, and Mathiowetz 2001):
consider the true model
Y=Xβ+ϵ
then
ˆβ=(˜X′˜X)−1˜X˜Y=(˜X′˜X)−1˜X′(˜Xβ−Uβ+v+ϵ)=β+(˜X′˜X)−1˜X′(−Uβ+v+ϵ)plimˆβ=β+plim(˜X′˜X)−1˜X′(−Uβ+v)=β+plim(˜X′˜X)−1˜X′W[−β1]
Since we collect the measurement errors in a matrix W=[U|v], then
(−Uβ+v)=W[−β1]
Hence, in general, biases in the coefficients β are regression coefficients from regressing the measurement errors on the mis-measured ˜X
Notes:
Instrumental Variable can help fix this problem
There can also be measurement error in dummy variables and you can still use Instrumental Variable to fix it.
36.1.1.3 Solution to Measurement Errors
36.1.1.3.1 Correlation
P(ρ|data)=P(data|ρ)P(ρ)P(data)Posterior Probability∝Likelihood×Prior Probability where
- ρ is a correlation coefficient
- P(data|ρ) is the likelihood function evaluated at ρ
- P(ρ) prior probability
- P(data) is the normalizing constant
With sample correlation coefficient r:
r=Sxy√SxxSyy Then the posterior density approximation of ρ is (Schisterman et al. 2003, 3)
P(ρ|x,y)∝P(ρ)(1−ρ2)(n−1)/2(1−ρ×r)n−(3/2)
where
- ρ=tanhξ where ξ∼N(z,1/n)
- r=tanhz
Then the posterior density follow a normal distribution where
Mean
μposterior=σ2posterior×(nprior×tanh−1rprior+nlikelihood×tanh−1rlikelihood)
variance
σ2posterior=1nprior+nLikelihood
To simplify the integration process, we choose prior that is
P(ρ)∝(1−ρ2)c where
- c is the weight the prior will have in estimation (i.e., c=0 if no prior info, hence P(ρ)∝1)
Example:
Current study: rxy=0.5,n=200
Previous study: rxy=0.2765,(n=50205)
Combining two, we have the posterior following a normal distribution with the variance of
σ2posterior=1nprior+nLikelihood=1200+50205=0.0000198393
Mean
μPosterior=σ2Posterior×(nprior×tanh−1rprior+nlikelihood×tanh−1rlikelihood)=0.0000198393×(50205×tanh−10.2765+200×tanh−10.5)=0.2849415
Hence, Posterior∼N(0.691,0.0009), which means the correlation coefficient is tanh(0.691)=0.598 and 95% CI is
μposterior±1.96×√σ2Posterior=0.2849415±1.96×(0.0000198393)1/2=(0.2762115,0.2936714)
Hence, the interval for posterior ρ is (0.2693952,0.2855105)
If future authors suspect that they have
- Large sampling variation
- Measurement error in either measures in the correlation, which attenuates the relationship between the two variables
Applying this Bayesian correction can give them a better estimate of the correlation between the two.
To implement this calculation in R, see below
n_new <- 200
r_new <- 0.5
alpha <- 0.05
update_correlation <- function(n_new, r_new, alpha) {
n_meta <- 50205
r_meta <- 0.2765
# Variance
var_xi <- 1 / (n_new + n_meta)
format(var_xi, scientific = FALSE)
# mean
mu_xi <- var_xi * (n_meta * atanh(r_meta) + n_new * (atanh(r_new)))
format(mu_xi, scientific = FALSE)
# confidence interval
upper_xi <- mu_xi + qnorm(1 - alpha / 2) * sqrt(var_xi)
lower_xi <- mu_xi - qnorm(1 - alpha / 2) * sqrt(var_xi)
# rho
mean_rho <- tanh(mu_xi)
upper_rho <- tanh(upper_xi)
lower_rho <- tanh(lower_xi)
# return a list
return(
list(
"mu_xi" = mu_xi,
"var_xi" = var_xi,
"upper_xi" = upper_xi,
"lower_xi" = lower_xi,
"mean_rho" = mean_rho,
"upper_rho" = upper_rho,
"lower_rho" = lower_rho
)
)
}
# Old confidence interval
r_new + qnorm(1 - alpha / 2) * sqrt(1/n_new)
#> [1] 0.6385904
r_new - qnorm(1 - alpha / 2) * sqrt(1/n_new)
#> [1] 0.3614096
testing = update_correlation(n_new = n_new, r_new = r_new, alpha = alpha)
# Updated rho
testing$mean_rho
#> [1] 0.2774723
# Updated confidence interval
testing$upper_rho
#> [1] 0.2855105
testing$lower_rho
#> [1] 0.269395236.1.2 Simultaneity
When independent variables (X’s) are jointly determined with the dependent variable Y, typically through an equilibrium mechanism, violates the second condition for causality (i.e., temporal order).
Examples: quantity and price by demand and supply, investment and productivity, sales and advertisement
General Simultaneous (Structural) Equations
Yi=β0+β1Xi+uiXi=α0+α1Yi+vi
Hence, the solutions are
Yi=β0+β1α01−α1β1+β1vi+ui1−α1β1Xi=α0+α1β01−α1β1+vi+α1ui1−α1β1
If we run only one regression, we will have biased estimators (because of simultaneity bias):
Cov(Xi,ui)=Cov(vi+α1ui1−α1β1,ui)=α11−α1β1Var(ui)
In an even more general model
{Yi=β0+β1Xi+β2Ti+uiXi=α0+α1Yi+α2Zi+vi
where
Xi,Yi are endogenous variables determined within the system
Ti,Zi are exogenous variables
Then, the reduced form of the model is
{Yi=β0+β1α01−α1β1+β1α21−α1β1Zi+β21−α1β1Ti+˜ui=B0+B1Zi+B2Ti+˜uiXi=α0+α1β01−α1β1+α21−α1β1Zi+α1β21−α1β1Ti+˜vi=A0+A1Zi+A2Ti+˜vi
Then, now we can get consistent estimates of the reduced form parameters
And to get the original parameter estimates
B1A1=β1B2(1−B1A2A1B2)=β2A2B2=α1A1(1−B1A2A1B2)=α2
Rules for Identification
Order Condition (necessary but not sufficient)
K−k≥m−1
where
M = number of endogenous variables in the model
K = number of exogenous variables int he model
m = number of endogenous variables in a given
k = is the number of exogenous variables in a given equation
This is actually the general framework for instrumental variables
36.1.3 Endogenous Treatment Solutions
Using the OLS estimates as a reference point
library(AER)
library(REndo)
set.seed(421)
data("CASchools")
school <- CASchools
school$stratio <- with(CASchools, students / teachers)
m1.ols <-
lm(read ~ stratio + english + lunch
+ grades + income + calworks + county,
data = school)
summary(m1.ols)$coefficients[1:7,]
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 683.45305948 9.56214469 71.4748711 3.011667e-218
#> stratio -0.30035544 0.25797023 -1.1643027 2.450536e-01
#> english -0.20550107 0.03765408 -5.4576041 8.871666e-08
#> lunch -0.38684059 0.03700982 -10.4523759 1.427370e-22
#> gradesKK-08 -1.91291321 1.35865394 -1.4079474 1.599886e-01
#> income 0.71615378 0.09832843 7.2832829 1.986712e-12
#> calworks -0.05273312 0.06154758 -0.8567863 3.921191e-0136.1.3.1 Instrumental Variable
[A3a] requires ϵi to be uncorrelated with xi
plim(ˆβOLS)=β+[E(x′ixi)]−1E(x′iϵi)
[A3a] is the weakest assumption needed for OLS to be consistent
A3 fails when xik is correlated with ϵi
- Omitted Variables Bias: ϵi includes any other factors that may influence the dependent variable (linearly)
- Simultaneity Demand and prices are simultaneously determined.
- Endogenous Sample Selection we did not have iid sample
- Measurement Error
Note
- Omitted Variable: an omitted variable is a variable, omitted from the model (but is in the ϵi) and unobserved has predictive power towards the outcome.
- Omitted Variable Bias: is the bias (and inconsistency when looking at large sample properties) of the OLS estimator when the omitted variable.
- We cam have both positive and negative selection bias (it depends on what our story is)
The structural equation is used to emphasize that we are interested understanding a causal relationship
yi1=β0+zi1β1+yi2β2+ϵi
where
- yit is the outcome variable (inherently correlated with ϵi)
- yi2 is the endogenous covariate (presumed to be correlated with ϵi)
- β1 represents the causal effect of yi2 on yi1
- zi1 is exogenous controls (uncorrelated with ϵi) (E(z′1iϵi)=0)
OLS is an inconsistent estimator of the causal effect β2
If there was no endogeneity
- E(y′i2ϵi)=0
- the exogenous variation in yi2 is what identifies the causal effect
If there is endogeneity
- Any wiggle in yi2 will shift simultaneously with ϵi
plim(ˆβOLS)=β+[E(x′ixi)]−1E(x′iϵi)
where
- β is the causal effect
- [E(x′ixi)]−1E(x′iϵi) is the endogenous effect
Hence ˆβOLS can be either more positive and negative than the true causal effect.
Motivation for Two Stage Least Squares (2SLS)
yi1=β0+zi1β1+yi2β2+ϵi
We want to understand how movement in yi2 effects movement in yi1, but whenever we move yi2, ϵi also moves.
Solution
We need a way to move yi2 independently of ϵi, then we can analyze the response in yi1 as a causal effect
-
Find an instrumental variable(s) zi2
- Instrument Relevance: when** zi2 moves then yi2 also moves
- Instrument Exogeneity: when zi2 moves then ϵi does not move.
zi2 is the exogenous variation that identifies the causal effect β2
Finding an Instrumental variable:
- Random Assignment: + Effect of class size on educational outcomes: instrument is initial random
- Relation’s Choice + Effect of Education on Fertility: instrument is parent’s educational level
- Eligibility + Trade-off between IRA and 401K retirement savings: instrument is 401k eligibility
Example
Return to College
education is correlated with ability - endogenous
-
Near 4year as an instrument
- Instrument Relevance: when near moves then education also moves
- Instrument Exogeneity: when near moves then ϵi does not move.
Other potential instruments; near a 2-year college. Parent’s Education. Owning Library Card
yi1=β0+zi1β1+yi2β2+ϵi
First Stage (Reduced Form) Equation:
yi2=π0+zi1π1+zi2π2+vi
where
- π0+zi1π1+zi2π2 is exogenous variation vi is endogenous variation
This is called a reduced form equation
Not interested in the causal interpretation of π1 or π2
A linear projection of zi1 and zi2 on yi2 (simple correlations)
The projections π1 and π2 guarantee that E(z′i1vi)=0 and E(z′i2vi)=0
Instrumental variable zi2
- Instrument Relevance: π2≠0
- Instrument Exogeneity: E(zi2ϵi)=0
Moving only the exogenous part of yi2 is moving
˜yi2=π0+zi1π1+zi2π2
two Stage Least Squares (2SLS)
yi1=β0+zi1β1+yi2β2+ϵi
yi2=π0+zi2π2+vi
Equivalently,
yi1=β0+zi1β1+˜yi2β2+uiwhere
- ˜yi2=π0+zi2π2
- ui=viβ2+ϵi
- A2 holds if the instrument is relevant π2≠0 + yi1=β0+zi1β1+(π0+zi1π1+zi2π2)β2+ui
- [A3a] holds if the instrument is exogenous E(zi2ϵi)=0
E(˜y′i2ui)=E((π0+zi1π1+zi2)(viβ2+ϵi))=E((π0+zi1π1+zi2)(ϵi))=E(ϵi)π0+E(ϵizi1)π1+E(ϵizi2)=0
Hence, (36.1) is consistent
The 2SLS Estimator
1. Estimate the first stage using OLS
yi2=π0+zi2π2+vi
and obtained estimated value ˆyi2
- Estimate the altered equation using OLS
yi1=β0+zi1β1+ˆyi2β2+ϵi
Properties of the 2SLS Estimator
- Under A1, A2, [A3a] (for zi1), A5 and if the instrument satisfies the following two conditions, + Instrument Relevance: π2≠0 + Instrument Exogeneity: E(z′i2ϵi)=0 then the 2SLS estimator is consistent
- Can handle more than one endogenous variable and more than one instrumental variable
yi1=β0+zi1β1+yi2β2+yi3β3+ϵiyi2=π0+zi1π1+zi2π2+zi3π3+zi4π4+vi2yi3=γ0+zi1γ1+zi2γ2+zi3γ3+zi4γ4+vi3
+ **IV estimator**: one endogenous variable with a single instrument
+ **2SLS estimator**: one endogenous variable with multiple instruments
+ **GMM estimator**: multiple endogenous variables with multiple instruments
-
Standard errors produced in the second step are not correct
- Because we do not know ˜y perfectly and need to estimate it in the firs step, we are introducing additional variation
- We did not have this problem with FGLS because “the first stage was orthogonal to the second stage.” This is generally not true for most multi-step procedure.
- If A4 does not hold, need to report robust standard errors.
-
2SLS is less efficient than OLS and will always have larger standard errors.
- First, Var(ui)=Var(viβ2+ϵi)>Var(ϵi)
- Second, ˆyi2 is generally highly collinear with zi1
- First, Var(ui)=Var(viβ2+ϵi)>Var(ϵi)
The number of instruments need to be at least as many or more the number of endogenous variables.
Note
- 2SLS can be combined with FGLS to make the estimator more efficient: You have the same first-stage, and in the second-stage, instead of using OLS, you can use FLGS with the weight matrix ˆw
- Generalized Method of Moments can be more efficient than 2SLS.
- In the second-stage of 2SLS, you can also use MLE, but then you are making assumption on the distribution of the outcome variable, the endogenous variable, and their relationship (joint distribution).
36.1.3.1.1 Testing Assumptions
Endogeneity Test: Is yi2 truly endogenous (i.e., can we just use OLS instead of 2SLS)?
Exogeneity (Cannot always test and when you can it might not be informative)
Relevancy (need to avoid “weak instruments”)
36.1.3.1.1.1 Endogeneity Test
2SLS is generally so inefficient that we may prefer OLS if there is not much endogeneity
Biased but inefficient vs efficient but biased
-
Want a sense of “how endogenous” yi2 is
- if “very” endogenous - should use 2SLS
- if not “very” endogenous - perhaps prefer OLS
Invalid Test of Endogeneity: yi2 is endogenous if it is correlated with ϵi,
ϵi=γ0+yi2γ1+errori
where γ1≠0 implies that there is endogeneity
- ϵi is not observed, but using the residuals
ei=γ0+yi2γ1+errori
is NOT a valid test of endogeneity + The OLS residual, e is mechanically uncorrelated with yi2 (by FOC for OLS) + In every situation, γ1 will be essentially 0 and you will never be able to reject the null of no endogeneity
Valid test of endogeneity
- If yi2 is not endogenous then ϵi and v are uncorrelated
yi1=β0+zi1β1+yi2β2+ϵiyi2=π0+zi1π1+zi2π2+vi
Variable Addition test: include the first stage residuals as an additional variable,
yi1=β0+zi1β1+yi2β2+ˆviθ+errori
Then the usual t-test of significance is a valid test to evaluate the following hypothesis. note this test requires your instrument to be valid instrument.
H0:θ=0 (not endogenous)H1:θ≠0 (endogenous)
36.1.3.1.1.2 Exogeneity
Why exogeneity matter?
E(z′i2ϵi)=0
- If [A3a] fails - 2SLS is also inconsistent
- If instrument is not exogenous, then we need to find a new one.
- Similar to Endogeneity Test, when there is a single instrument
ei=γ0+zi2γ1+erroriH0:γ1=0
is NOT a valid test of endogeneity
- the OLS residual, e is mechanically uncorrelated with zi2: ˆγ1 will be essentially 0 and you will never be able to determine if the instrument is endogenous.
Solution
Testing Instrumental Exogeneity in an Over-identified Model
-
When there is more than one exogenous instrument (per endogenous variable), we can test for instrument exogeneity.
When we have multiple instruments, the model is said to be over-identified.
Could estimate the same model several ways (i.e., can identify/ estimate β1 more than one way)
Idea behind the test: if the controls and instruments are truly exogenous then OLS estimation of the following regression,
ϵi=γ0+zi1γ1+zi2γ2+errori
should have a very low R2
- if the model is just identified (one instrument per endogenous variable) then the R2=0
Steps:
Estimate the structural equation by 2SLS (using all available instruments) and obtain the residuals e
Regress e on all controls and instruments and obtain the R2
-
Under the null hypothesis (all IV’s are uncorrelated), nR2∼χ2(q), where q is the number of instrumental variables minus the number of endogenous variables
- if the model is just identified (one instrument per endogenous variable) then q = 0, and the distribution under the null collapses.
low p-value means you reject the null of exogenous instruments. Hence you would like to have high p-value in this test.
Pitfalls for the Overid test
-
the overid test is essentially compiling the following information.
- Conditional on first instrument being exogenous is the other instrument exogenous?
- Conditional on the other instrument being exogenous, is the first instrument exogenous?
If all instruments are endogenous than neither test will be valid
really only useful if one instrument is thought to be truly exogenous (randomly assigned). even f you do reject the null, the test does not tell you which instrument is exogenous and which is endogenous.
| Result | Implication |
|---|---|
| reject the null | you can be pretty sure there is an endogenous instrument, but don’t know which one. |
| fail to reject | could be either (1) they are both exogenous, (2) they are both endogenous. |
36.1.3.1.1.3 Relevancy
Why Relevance matter?
π2≠0
-
used to show A2 holds
If π2=0 (instrument is not relevant) then A2 fails - perfect multicollinearity
If π2 is close to 0 (weak instrument) then there is near perfect multicollinearity - 2SLS is highly inefficient (Large standard errors).
-
A weak instrument will exacerbate any inconsistency due to an instrument being (even slightly) endogenous.
- In the simple case with no controls and a single endogenous variable and single instrumental variable,
plim(ˆβ22SLS)=β2+E(zi2ϵi)E(zi2yi2)
Testing Weak Instruments
can use t-test (or F-test for over-identified models) in the first stage to determine if there is a weak instrument problem.
-
J. Stock and Yogo (2005): a statistical rejection of the null hypothesis in the first stage at the 5% (or even 1%) level is not enough to insure the instrument is not weak
- Rule of Thumb: need a F-stat of at least 10 (or a t-stat of at least 3.2) to reject the null hypothesis that the instrument is weak.
Summary of the 2SLS Estimator
yi1=β0+zi1β1+yi2β2+ϵiyi2=π0+zi1π1+zi2π2+vi
- when [A3a] does not hold
E(y′i2ϵi)≠0
Then the OLS estimator is no longer unbiased or consistent.
If we have valid instruments zi2
-
Relevancy (need to avoid “weak instruments”): π2≠0 Then the 2SLS estimator is consistent under A1, A2, [A5a], and the above two conditions.
yi1=β0+zi1β1+yi2β2+ϵiyi2=π0+zi1π1+zi2π2+vi
- When [A3a] does hold
E(y′i2ϵi)=0
and we have valid instruments, then both the OLS and 2SLS estimators are consistent.
- The OLS estimator is always more efficient
- can use the variable addition test to determine if 2SLS is need (A3a does hold) or if OLS is valid (A3a does not hold)
Sometimes we can test the assumption for instrument to be valid:
- Exogeneity : Only table when there are more instruments than endogenous variables.
- Relevancy (need to avoid “weak instruments”): Always testable, need the F-stat to be greater than 10 to rule out a weak instrument
Application
Expenditure as observed instrument
m2.2sls <-
ivreg(
read ~ stratio + english + lunch
+ grades + income + calworks + county |
expenditure + english + lunch
+ grades + income + calworks + county ,
data = school
)
summary(m2.2sls)$coefficients[1:7,]
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 700.47891593 13.58064436 51.5792106 8.950497e-171
#> stratio -1.13674002 0.53533638 -2.1234126 3.438427e-02
#> english -0.21396934 0.03847833 -5.5607753 5.162571e-08
#> lunch -0.39384225 0.03773637 -10.4366757 1.621794e-22
#> gradesKK-08 -1.89227865 1.37791820 -1.3732881 1.704966e-01
#> income 0.62487986 0.11199008 5.5797785 4.668490e-08
#> calworks -0.04950501 0.06244410 -0.7927892 4.284101e-0136.1.3.1.2 Checklist
- Regress the dependent variable on the instrument (reduced form). Since under OLS, we have unbiased estimate, the coefficient estimate should be significant (make sure the sign makes sense)
- Report F-stat on the excluded instruments. F-stat < 10 means you have a weak instrument (J. H. Stock, Wright, and Yogo 2002).
- Present R2 before and after including the instrument (Rossi 2014)
- For models with multiple instrument, present firs-t and second-stage result for each instrument separately. Overid test should be conducted (e.g., Sargan-Hansen J)
- Hausman test between OLS and 2SLS (don’t confuse this test for evidence that endogeneity is irrelevant - under invalid IV, the test is useless)
- Compare the 2SLS with the limited information ML. If they are different, you have evidence for weak instruments.
36.1.3.2 Good Instruments
Exogeneity and Relevancy are necessary but not sufficient for IV to produce consistent estimates.
Without theory or possible explanation, you can always create a new variable that is correlated with X and uncorrelated with ϵ
For example, we want to estimate the effect of price on quantity (Reiss 2011, 960)
Q=β1P+β2X+ϵP=π1X+η
where ϵ and η are jointly determined, X⊥ϵ,η
Without theory, we can just create a new variable Z=X+u where E(u)=0;u⊥X,ϵ,η
Then, Z satisfied both conditions:
Relevancy: X correlates P → Z correlates P
Exogeneity: u⊥ϵ (random noise)
But obviously, it’s not a valid instrument (intuitively). But theoretically, relevance and exogeneity are not sufficient to identify β because of unsatisfied rank condition for identification.
Moreover, the functional form of the instrument also plays a role when choosing a good instrument. Hence, we always need to check for the robustness of our instrument.
IV methods even with valid instruments can still have poor sampling properties (finite sample bias, large sampling errors) (Rossi 2014)
When you have a weak instrument, it’s important to report it appropriately. This problem will be exacerbated if you have multiple instruments (Larcker and Rusticus 2010).
36.1.3.2.1 Lagged dependent variable
In time series data sets, we can use lagged dependent variable as an instrument because it is not influenced by current shocks. For example, Chetty, Friedman, and Rockoff (2014) used lagged dependent variable in econ.
36.1.3.2.2 Lagged explanatory variable
-
Common practice in applied economics: Replace suspected simultaneously determined explanatory variable with its lagged value Bellemare, Masaki, and Pepinsky (2017).
This practice does not avoid simultaneity bias.
Estimates using this method are still inconsistent.
Hypothesis testing becomes invalid under this approach.
Lagging variables changes how endogeneity bias operates, adding a “no dynamics among unobservables” assumption to the “selection on observables” assumption.
-
Key conditions for appropriate use (Bellemare, Masaki, and Pepinsky 2017):
-
Under unobserved confounding:
- No dynamics among unobservables.
- The lagged variable X is a stationary autoregressive process.
-
Under no unobserved confounding:
- No reverse causality; the causal effect operates with a one-period lag (Xt−1→Y, Xt↛).
- Reverse causality is contemporaneous, with a one-period lag effect.
- Reverse causality is contemporaneous; no dynamics in Y, but dynamics exist in X (X_{t-1} \to X).
-
Under unobserved confounding:
-
Alternative approach: Use lagged values of the endogenous variable in IV estimation. However, IV estimation is only effective if (Reed 2015):
Lagged values do not belong in the estimating equation.
Lagged values are sufficiently correlated with the simultaneously determined explanatory variable.
Lagged IVs help mitigate endogeneity if they only violate the independence assumption. However, if lagged IVs violate both the independence assumption and exclusion restriction, they may aggravate endogeneity (Yu Wang and Bellemare 2019).
36.1.3.3 Internal instrumental variable
(also known as instrument free methods). This section is based on Raluca Gui’s guide
alternative to external instrumental variable approaches
All approaches here assume a continuous dependent variable
36.1.3.3.1 Non-hierarchical Data (Cross-classified)
Y_t = \beta_0 + \beta_1 P_t + \beta_2 X_t + \epsilon_t
where
- t = 1, .., T (indexes either time or cross-sectional units)
- Y_t is a k \times 1 response variable
- X_t is a k \times n exogenous regressor
- P_t is a k \times 1 continuous endogenous regressor
- \epsilon_t is a structural error term with \mu_\epsilon =0 and E(\epsilon^2) = \sigma^2
- \beta are model parameters
The endogeneity problem arises from the correlation of P_t and \epsilon_t:
P_t = \gamma Z_t + v_t
where
- Z_t is a l \times 1 vector of internal instrumental variables
- ν_t is a random error with \mu_{v_t}, E(v^2) = \sigma^2_v, E(\epsilon v) = \sigma_{\epsilon v}
- Z_t is assumed to be stochastic with distribution G
- ν_t is assumed to have density h(·)
36.1.3.3.1.1 Latent Instrumental Variable
assume Z_t (unobserved) to be uncorrelated with \epsilon_t, which is similar to Instrumental Variable. Hence, Z_t and ν_t can’t be identified without distributional assumptions
The distributions of Z_t and ν_t need to be specified such that:
- endogeneity of P_t is corrected
- the distribution of P_t is empirically close to the integral that expresses the amount of overlap of Z as it is shifted over ν (= the convolution between Z_t and ν_t).
When the density h(·) = Normal, then G cannot be normal because the parameters would not be identified (Ebbes et al. 2005) .
Hence,
- in the LIV model the distribution of Z_t is discrete
- in the Higher Moments Method and Joint Estimation Using Copula methods, the distribution of Z_t is taken to be skewed.
Z_t are assumed unobserved, discrete and exogenous, with
- an unknown number of groups m
- \gamma is a vector of group means.
Identification of the parameters relies on the distributional assumptions of
- P_t: a non-Gaussian distribution
- Z_t discrete with m \ge 2
Note:
- If Z_t is continuous, the model is unidentified
- If P_t \sim N, you have inefficient estimates.
m3.liv <- latentIV(read ~ stratio, data = school)
summary(m3.liv)$coefficients[1:7, ]
#> Estimate Std. Error z-score Pr(>|z|)
#> (Intercept) 6.996014e+02 2.686165e+02 2.604462e+00 9.529035e-03
#> stratio -2.272673e+00 1.367747e+01 -1.661618e-01 8.681097e-01
#> pi1 -4.896363e+01 NaN NaN NaN
#> pi2 1.963920e+01 9.225351e-02 2.128830e+02 0.000000e+00
#> theta5 6.939432e-152 3.143456e-161 2.207581e+09 0.000000e+00
#> theta6 3.787512e+02 4.249436e+01 8.912976e+00 1.541010e-17
#> theta7 -1.227543e+00 4.885237e+01 -2.512761e-02 9.799651e-01it will return a coefficient very different from the other methods since there is only one endogenous variable.
36.1.3.3.1.2 Joint Estimation Using Copula
assume Z_t (unobserved) to be uncorrelated with \epsilon_t, which is similar to Instrumental Variable. Hence, Z_t and ν_t can’t be identified without distributional assumptions
(S. Park and Gupta 2012) allows joint estimation of the continuous P_t and \epsilon_t using Gaussian copulas, where a copula is a function that maps several conditional distribution functions (CDF) into their joint CDF).
The underlying idea is that using information contained in the observed data, one selects marginal distributions for P_t and \epsilon_t. Then, the copula model constructs a flexible multivariate joint distribution that allows a wide range of correlations between the two marginals.
The method allows both continuous and discrete P_t.
In the special case of one continuous P_t, estimation is based on MLE
Otherwise, based on Gaussian copulas, augmented OLS estimation is used.
Assumptions:
skewed P_t
the recovery of the correct parameter estimates
\epsilon_t \sim normal marginal distribution. The marginal distribution of P_t is obtained using the Epanechnikov kernel density estimator
\hat{h}_p = \frac{1}{T . b} \sum_{t=1}^TK(\frac{p - P_t}{b}) whereP_t = endogenous variables
K(x) = 0.75(1-x^2)I(||x||\le 1)
-
b=0.9T^{-1/5}\times min(s, IQR/1.34)
- IQR = interquartile range
- s = sample standard deviation
- T = n of time periods observed in the data
In augmented OLS and MLE, the inference procedure occurs in two stages:
(1): the empirical distribution of P_t is computed
(2) used in it constructing the likelihood function)
Hence, the standard errors would not be correct.
So we use the sampling distributions (from bootstrapping) to get standard errors and the variance-covariance matrix. Since the distribution of the bootstrapped parameters is highly skewed, we report the percentile confidence intervals is preferable.
set.seed(110)
m4.cc <-
copulaCorrection(
read ~ stratio + english + lunch + calworks +
grades + income + county |
continuous(stratio),
data = school,
optimx.args = list(method = c("Nelder-Mead"),
itnmax = 60000),
num.boots = 2,
verbose = FALSE
)
summary(m4.cc)$coefficients[1:7,]
#> Point Estimate Boots SE Lower Boots CI (95%)
#> (Intercept) 682.38500189 2.438874461 NA
#> stratio -0.31091374 0.022332810 NA
#> english -0.19648765 0.026136685 NA
#> lunch -0.38439247 0.032293205 NA
#> calworks -0.04363619 0.009027372 NA
#> gradesKK-08 -1.97957885 0.138420392 NA
#> income 0.77449995 0.033266598 NA
#> Upper Boots CI (95%)
#> (Intercept) NA
#> stratio NA
#> english NA
#> lunch NA
#> calworks NA
#> gradesKK-08 NA
#> income NAwe run this model with only one endogenous continuous regressor (stratio). Sometimes, the code will not converge, in which case you can use different
- optimization algorithm
- starting values
- maximum number of iterations
36.1.3.3.1.3 Higher Moments Method
suggested by (Lewbel 1997) to identify \epsilon_t caused by measurement error.
Identification is achieved by using third moments of the data, with no restrictions on the distribution of \epsilon_t
The following instruments can be used with 2SLS estimation to obtain consistent estimates:
\begin{aligned} q_{1t} &= (G_t - \bar{G}) \\ q_{2t} &= (G_t - \bar{G})(P_t - \bar{P}) \\ q_{3t} &= (G_t - \bar{G})(Y_t - \bar{Y})\\ q_{4t} &= (Y_t - \bar{Y})(P_t - \bar{P}) \\ q_{5t} &= (P_t - \bar{P})^2 \\ q_{6t} &= (Y_t - \bar{Y})^2 \\ \end{aligned}
where
- G_t = G(X_t) for any given function G that has finite third own and cross moments
- X = exogenous variable
q_{5t}, q_{6t} can be used only when the measurement and \epsilon_t are symmetrically distributed. The rest of the instruments does not require any distributional assumptions for \epsilon_t.
Since the regressors G(X) = X are included as instruments, G(X) can’t be a linear function of X in q_{1t}
Since this method has very strong assumptions, Higher Moments Method should only be used in case of overidentification
set.seed(111)
m5.hetEr <-
hetErrorsIV(
read ~ stratio + english + lunch + calworks + income +
grades + county |
stratio | IIV(income, english),
data = school
)
summary(m5.hetEr)$coefficients[1:7,]
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 662.78791557 27.90173069 23.7543657 2.380436e-76
#> stratio 0.71480686 1.31077325 0.5453322 5.858545e-01
#> english -0.19522271 0.04057527 -4.8113717 2.188618e-06
#> lunch -0.37834232 0.03927793 -9.6324402 9.760809e-20
#> calworks -0.05665126 0.06302095 -0.8989273 3.692776e-01
#> income 0.82693755 0.17236557 4.7975797 2.335271e-06
#> gradesKK-08 -1.93795843 1.38723186 -1.3969968 1.632541e-01recommend using this approach to create additional instruments to use with external ones for better efficiency.
36.1.3.3.1.4 Heteroskedastic Error Approach
- using means of variables that are uncorrelated with the product of heteroskedastic errors to identify structural parameters.
- This method can be use either when you don’t have external instruments or you want to use additional instruments to improve the efficiency of the IV estimator (Lewbel 2012)
- The instruments are constructed as simple functions of data
- Model’s assumptions:
\begin{aligned} E(X \epsilon) &= 0 \\ E(X v ) &= 0 \\ cov(Z, \epsilon v) &= 0 \\ cov(Z, v^2) &\neq 0 \text{ (for identification)} \end{aligned}
Structural parameters are identified by 2SLS regression of Y on X and P, using X and [Z − E(Z)]ν as instruments.
\text{instrument's strength} \propto cov((Z-\bar{Z})v,v)
where cov((Z-\bar{Z})v,v) is the degree of heteroskedasticity of ν with respect to Z (Lewbel 2012), which can be empirically tested.
If it is zero or close to zero (i.e.,the instrument is weak), you might have imprecise estimates, with large standard errors.
- Under homoskedasticity, the parameters of the model are unidentified.
- Under heteroskedasticity related to at least some elements of X, the parameters of the model are identified.
36.1.3.3.2 Hierarchical Data
Multiple independent assumptions involving various random components at different levels mean that any moderate correlation between some predictors and a random component or error term can result in a significant bias of the coefficients and of the variance components. (J.-S. Kim and Frees 2007) proposed a generalized method of moments which uses both, the between and within variations of the exogenous variables, but only assumes the within variation of the variables to be endogenous.
Assumptions
- the errors at each level \sim iid N
- the slope variables are exogenous
- the level-1 \epsilon \perp X, P. If this is not the case, additional, external instruments are necessary
Hierarchical Model
\begin{aligned} Y_{cst} &= Z_{cst}^1 \beta_{cs}^1 + X_{cst}^1 \beta_1 + \epsilon_{cst}^1 \\ \beta^1_{cs} &= Z_{cs}^2 \beta_{c}^2 + X_{cst}^2 \beta_2 + \epsilon_{cst}^2 \\ \beta^2_{c} &= X^3_c \beta_3 + \epsilon_c^3 \end{aligned}
Bias could stem from:
- errors at the higher two levels (\epsilon_c^3,\epsilon_{cst}^2) are correlated with some of the regressors
- only third level errors (\epsilon_c^3) are correlated with some of the regressors
(J.-S. Kim and Frees 2007) proposed
- When all variables are assumed exogenous, the proposed estimator equals the random effects estimator
- When all variables are assumed endogenous, it equals the fixed effects estimator
- also use omitted variable test (based on the Hausman-test (J. A. Hausman 1978) for panel data), which allows the comparison of a robust estimator and an estimator that is efficient under the null hypothesis of no omitted variables or the comparison of two robust estimators at different levels.
# function 'cholmod_factor_ldetA' not provided by package 'Matrix'
set.seed(113)
school$gr08 <- school$grades == "KK-06"
m7.multilevel <-
multilevelIV(read ~ stratio + english + lunch + income + gr08 +
calworks + (1 | county) | endo(stratio),
data = school)
summary(m7.multilevel)$coefficients[1:7,]Another example using simulated data
- level-1 regressors: X_{11}, X_{12}, X_{13}, X_{14}, X_{15}, where X_{15} is correlated with the level-2 error (i.e., endogenous).
- level-2 regressors: X_{21}, X_{22}, X_{23}, X_{24}
- level-3 regressors: X_{31}, X_{32}, X_{33}
We estimate a three-level model with X15 assumed endogenous. Having a three-level hierarchy, multilevelIV() returns five estimators, from the most robust to omitted variables (FE_L2), to the most efficient (REF) (i.e. lowest mean squared error).
- The random effects estimator (REF) is efficient assuming no omitted variables
- The fixed effects estimator (FE) is unbiased and asymptotically normal even in the presence of omitted variables.
- Because of the efficiency, the random effects estimator is preferable if you think there is no omitted. variables
- The robust estimator would be preferable if you think there is omitted variables.
# function 'cholmod_factor_ldetA' not provided by package 'Matrix'’
data(dataMultilevelIV)
set.seed(114)
formula1 <-
y ~ X11 + X12 + X13 + X14 + X15 + X21 + X22 + X23 + X24 +
X31 + X32 + X33 + (1 | CID) + (1 | SID) | endo(X15)
m8.multilevel <-
multilevelIV(formula = formula1, data = dataMultilevelIV)
coef(m8.multilevel)
summary(m8.multilevel, "REF")True \beta_{X_{15}} =-1. We can see that some estimators are bias because X_{15} is correlated with the level-two error, to which only FE_L2 and GMM_L2 are robust
To select the appropriate estimator, we use the omitted variable test.
In a three-level setting, we can have different estimator comparisons:
- Fixed effects vs. random effects estimators: Test for omitted level-two and level-three omitted effects, simultaneously, one compares FE_L2 to REF. But we will not know at which omitted variables exist.
- Fixed effects vs. GMM estimators: Once the existence of omitted effects is established but not sure at which level, we test for level-2 omitted effects by comparing FE_L2 vs GMM_L3. If you reject the null, the omitted variables are at level-2 The same is accomplished by testing FE_L2 vs. GMM_L2, since the latter is consistent only if there are no omitted effects at level-2.
- Fixed effects vs. fixed effects estimators: We can test for omitted level-2 effects, while allowing for omitted level-3 effects by comparing FE_L2 vs. FE_L3 since FE_L2 is robust against both level-2 and level-3 omitted effects while FE_L3 is only robust to level-3 omitted variables.
Summary, use the omitted variable test comparing REF vs. FE_L2 first.
If the null hypothesis is rejected, then there are omitted variables either at level-2 or level-3
-
Next, test whether there are level-2 omitted effects, since testing for omitted level three effects relies on the assumption there are no level-two omitted effects. You can use any of these pair of comparisons:
FE_L2 vs. FE_L3FE_L2 vs. GMM_L2
-
If no omitted variables at level-2 are found, test for omitted level-3 effects by comparing either
-
FE_L3vs.GMM_L3 -
GMM_L2vs.GMM_L3
-
summary(m8.multilevel, "REF")
# compare REF with all the other estimators. Testing REF (the most efficient estimator) against FE_L2 (the most robust estimator), equivalently we are testing simultaneously for level-2 and level-3 omitted effects. Since the null hypothesis is rejected (p = 0.000139), there is bias in the random effects estimator.
To test for level-2 omitted effects (regardless of level-3 omitted effects), we compare FE_L2 versus FE_L3
summary(m8.multilevel,"FE_L2")The null hypothesis of no omitted level-2 effects is rejected (p = 3.92e − 05). Hence, there are omitted effects at level-two. We should use FE_L2 which is consistent with the underlying data that we generated (level-2 error correlated with X_15, which leads to biased FE_L3 coefficients.
The omitted variable test between FE_L2 and GMM_L2 should reject the null hypothesis of no omitted level-2 effects (p-value is 0).
If we assume an endogenous variable as exogenous, the RE and GMM estimators will be biased because of the wrong set of internal instrumental variables. To increase our confidence, we should compare the omitted variable tests when the variable is considered endogenous vs. exogenous to get a sense whether the variable is truly endogenous.
36.1.3.4 Proxy Variables
Can be in place of the omitted variable
will not be able to estimate the effect of the omitted variable
will be able to reduce some endogeneity caused bye the omitted variable
but it can have Measurement Error. Hence, you have to be extremely careful when using proxies.
Criteria for a proxy variable:
- The proxy is correlated with the omitted variable.
- Having the omitted variable in the regression will solve the problem of endogeneity
- The variation of the omitted variable unexplained by the proxy is uncorrelated with all independent variables, including the proxy.
IQ test can be a proxy for ability in the regression between wage explained education.
For the third requirement
ability = \gamma_0 + \gamma_1 IQ + \epsilon
where \epsilon is uncorrelated with education and IQ test.
36.2 Endogenous Sample Selection
Endogenous sample selection arises in observational or non-experimental research whenever the inclusion of observations (or assignment to treatment) is not random, and the same unobservable factors influencing selection also affect the outcome of interest. This scenario leads to biased and inconsistent estimates of causal parameters (e.g., [Average Treatment Effects]) if not properly addressed.
This problem was first formalized in the econometric literature by J. Heckman (1974) J. J. Heckman (1976b) J. J. Heckman (1979), whose work addressed the issue in the context of labor force participation among women. Later, Amemiya (1984) generalize the method. Now, it has since been applied widely across social sciences, epidemiology, marketing, and finance.
Endogenous sample selection is often conflated with general selection bias, but it is important to understand that sample selection refers specifically to the inclusion of observations into the estimation sample, not just to assignment into treatment (i.e., selection bias).
This problem comes in many names such as self-selection problem, incidental truncation, or omitted variable (i.e., the omitted variable is how people were selected into the sample). Some disciplines consider nonresponse/selection bias as sample selection:
- When unobservable factors that affect who is in the sample are independent of unobservable factors that affect the outcome, the sample selection is not endogenous. Hence, the sample selection is ignorable and estimator that ignores sample selection is still consistent.
- When the unobservable factors that affect who is included in the sample are correlated with the unobservable factors that affect the outcome, the sample selection is endogenous and not ignorable, because estimators that ignore endogenous sample selection are not consistent (we don’t know which part of the observable outcome is related to the causal relationship and which part is due to different people were selected for the treatment and control groups).
Many evaluation studies use observational data, and in such data:
- Participants are not randomly assigned.
- Treatment or exposure is determined by individual or institutional choices.
- Counterfactual outcomes are not observed.
- The treatment indicator is often endogenous.
Some notable terminologies include:
-
Truncation: Occurs when data are collected only from a restricted subpopulation based on the value of a variable.
- Left truncation: Values below a threshold are excluded (e.g., only high-income individuals are surveyed).
- Right truncation: Values above a threshold are excluded.
-
Censoring: Occurs when the variable is observed but coarsened beyond a threshold.
- E.g., incomes below a certain level are coded as zero; arrest counts above a threshold are top-coded.
-
Incidental Truncation: Refers to selection based on a latent variable (e.g., employment decisions), not directly observed. This is what makes Heckman’s model distinct.
- Also called non-random sample selection.
- The error in the outcome equation is correlated with the selection indicator.
Researchers often categorize self-selection into:
-
Negative (Mitigation-Based) Selection: Individuals select into a treatment or sample to address an existing problem, so they start off with worse potential outcomes.
- Bias direction: Underestimates true treatment effects (makes the treatment look less effective than it is).
- Individuals select into treatment to combat a problem they already face.
-
Examples:
- People at high risk of severe illness (e.g., elderly or immunocompromised individuals) are more likely to get vaccinated. If we compare vaccinated vs. unvaccinated individuals without adjusting for risk factors, we might mistakenly conclude that vaccines are ineffective simply because vaccinated individuals had worse initial health conditions.
- Evaluating the effect of job training programs—unemployed individuals with the greatest difficulty finding jobs are most likely to enroll, leading to underestimated program benefits.
-
Positive (Preference-Based) Selection: Individuals select into a treatment or sample because they have advantageous traits, preferences, or resources. Hence, those who take treatment are systematically better off compared to those who do not.
- Bias direction: Overestimates true treatment effects (makes the treatment look more effective than it is).
- Individuals select into treatment because they inherently prefer it, rather than because of an underlying problem.
-
Examples:
- People who are health-conscious and physically active are more likely to join a fitness program. If we compare fitness program participants to non-participants, we might falsely attribute their better health outcomes to the program, when in reality, their pre-existing lifestyle contributed to their improved health.
- Evaluating the effect of private school education—students who attend private schools often come from wealthier families with greater academic support, making it difficult to isolate the true impact of the school itself.
Both forms of selection reflect correlation between unobservables (driving selection) and potential outcomes—the hallmark of endogenous selection bias.
Some seminal applied works in this area include:
- Labor Force Participation (J. Heckman 1974)
- Wages are observed only for women who choose to work.
- Unobservable preferences (reservation wages) drive participation.
- Ignoring this leads to biased estimates of the returns to education.
- Union Membership (Lewis 1986)
- Wages differ between union and non-union workers.
- But union membership is not exogenous—workers choose to join based on anticipated benefits.
- Naïve OLS yields biased estimates of union premium.
- Comparing income of college graduates vs. non-graduates.
- Attending college is a choice based on expected gains, ability, or family background.
- A treatment effect model (described next) is more appropriate here.
36.2.1 Unifying Model Frameworks
Though often conflated, there are several overlapping models to address endogenous selection:
- Sample Selection Model (J. J. Heckman 1979): Outcome is unobserved if an agent is not selected into the sample.
- Treatment Effect Model: Outcome is observed for both groups (treated vs. untreated), but treatment assignment is endogenous.
- Heckman-Type / Control Function Approaches: Decompose the endogenous regressor or incorporate a correction term (Inverse Mills Ratio or residual) to control for endogeneity.
All revolve around the challenge: unobserved factors affect both who is included (or treated) and outcomes.
To formalize the problem, we consider the outcome and selection equations. Let:
- y_i: observed outcome (e.g., wage)
- x_i: covariates affecting outcome
- z_i: covariates affecting selection
- w_i: binary indicator for selection into the sample (e.g., employment)
36.2.1.1 Sample Selection Model
We begin with an outcome equation, which describes the variable of interest y_i. However, we only observe y_i if a certain selection mechanism indicates it is part of the sample. That mechanism is captured by a binary indicator w_i = 1. Formally, the observed outcome equation is:
\begin{aligned} y_i &= x_i' \beta + \varepsilon_i, \quad &\text{(Observed only if } w_i = 1\text{)}, \\ \varepsilon_i &\sim N(0, \sigma_\varepsilon^2). \end{aligned}
Here, x_i is a vector of explanatory variables (or covariates) that explain y_i. The noise term \varepsilon_i is assumed to be normally distributed with mean zero and variance \sigma_\varepsilon^2. However, because we only see y_i for those cases in which w_i = 1, we must account for how the selection occurs.
Next, we specify the selection equation via a latent index model. Let w_i^* be an unobserved latent variable:
\begin{aligned} w_i^* &= z_i' \gamma + u_i, \\ w_i &= \begin{cases} 1 & \text{if } w_i^* > 0, \\ 0 & \text{otherwise}. \end{cases} \end{aligned}
Here, z_i is a vector of variables that influence whether or not y_i is observed. In practice, z_i may overlap with x_i, but can also include variables not in the outcome equation. For identification, we normalize \mathrm{Var}(u_i) = 1. This is analogous to the probit model’s standard normalization.
Because w_i = 1 exactly when w_i^* > 0, this event occurs if u_i \ge -\,z_i' \gamma. Therefore,
\begin{aligned} P(w_i = 1) &= P\bigl(u_i \ge -z_i' \gamma\bigr),\\ &= 1 - \Phi\bigl(-z_i'\gamma\bigr), \\ &= \Phi\bigl(z_i'\gamma\bigr), \end{aligned}
where we use the symmetry of the standard normal distribution.
We assume (\varepsilon_i, u_i) are jointly normally distributed with correlation \rho. In other words,
\begin{pmatrix} \varepsilon_i \\ u_i \end{pmatrix} \;\sim\; \mathcal{N} \!\Biggl( \begin{pmatrix} 0 \\ 0 \end{pmatrix}, \begin{pmatrix} \sigma^2_\varepsilon & \rho \,\sigma_\varepsilon \\ \rho \,\sigma_\varepsilon & 1 \end{pmatrix} \Biggr).
- If \rho = 0, the selection is exogenous: whether y_i is observed is unrelated to unobserved determinants of y_i.
- If \rho \neq 0, sample selection is endogenous. Failing to model this selection mechanism leads to biased estimates of \beta.
Interpreting \rho
- \rho > 0: Individuals with higher unobserved components in \varepsilon_i (and thus typically larger y_i) are more likely to appear in the sample. (Positive selection)
- \rho < 0: Individuals with higher unobserved components in \varepsilon_i are less likely to appear. (Negative selection)
- \rho = 0: No endogenous selection. Observed outcomes are effectively random with respect to the unobserved part of y_i.
In empirical practice, \rho signals the direction of bias one might expect if the selection process is ignored.
Often, it is helpful to visualize how part of the distribution of u_i (the error in the selection equation) is truncated based on the threshold w_i^*>0. Below is a notional R code snippet that draws a normal density and shades the region where u_i > -z_i'\gamma.
x = seq(-3, 3, length = 200)
y = dnorm(x, mean = 0, sd = 1)
plot(x,
y,
type = "l",
main = bquote("Probabibility distribution of" ~ u[i]))
x_shaded = seq(0.3, 3, length = 100)
y_shaded = dnorm(x_shaded, 0, 1)
polygon(c(0.3, x_shaded, 3), c(0, y_shaded, 0), col = "gray")
text(1, 0.1, expression(1 - Phi(-z[i] * gamma)))
arrows(-0.5, 0.1, 0.3, 0, length = 0.15)
text(-0.5, 0.12, expression(-z[i] * gamma))
legend("topright",
"Gray = Prob of Observed",
pch = 1,
inset = 0.02)
In this figure, the gray‐shaded area represents u_i > -z_i'\gamma. Observations in that range are included in the sample. If \rho\neq 0, then the unobserved factors that drive u_i also affect \varepsilon_i, causing a non‐representative sample of \varepsilon_i.
A core insight of the Heckman model is the conditional expectation of y_i given w_i=1:
E\bigl(y_i \mid w_i = 1\bigr) \;=\; E\bigl(y_i \mid w_i^*>0\bigr) \;=\; E\bigl(x_i'\beta + \varepsilon_i \mid u_i > -z_i'\gamma\bigr).
Since x_i'\beta is nonrandom (conditional on x_i), we get
E\bigl(y_i \mid w_i=1\bigr) = x_i'\beta + E\bigl(\varepsilon_i \mid u_i > -z_i'\gamma\bigr).
From bivariate normal properties:
\varepsilon_i \,\bigl\lvert\, u_i=a \;\sim\; N\!\Bigl(\rho\,\sigma_{\varepsilon}\cdot a,\; (1-\rho^2)\,\sigma_{\varepsilon}^2\Bigr).
Thus,
E\bigl(\varepsilon_i \mid u_i > -z_i'\gamma\bigr) =\; \rho\,\sigma_{\varepsilon}\, E\bigl(u_i \mid u_i > -z_i'\gamma\bigr).
If U\sim N(0,1), then
E(U \mid U>a) = \frac{\phi(a)}{1-\Phi(a)} = \frac{\phi(a)}{\Phi(-a)}, where \phi is the standard normal pdf, \Phi is the cdf. By symmetry, \phi(-a)=\phi(a) and \Phi(-a)=1-\Phi(a). Letting a = -\,z_i'\gamma yields
E\bigl(U \mid U > -z_i'\gamma\bigr) = \frac{\phi(-z_i'\gamma)}{1-\Phi(-z_i'\gamma)} = \frac{\phi(z_i'\gamma)}{\Phi(z_i'\gamma)}.
Define the Inverse Mills Ratio (IMR) as
\lambda(x) = \frac{\phi(x)}{\Phi(x)}.
Hence,
E\bigl(\varepsilon_i \mid u_i > -z_i'\gamma\bigr) = \rho\,\sigma_{\varepsilon}\,\lambda\bigl(z_i'\gamma\bigr), and therefore
\boxed{ E\bigl(y_i \mid w_i=1\bigr) = x_i'\beta \;+\; \rho\,\sigma_{\varepsilon}\, \frac{\phi\bigl(z_i'\gamma\bigr)}{\Phi\bigl(z_i'\gamma\bigr)}. }
This extra term is the so‐called Heckman correction.
The IMR appears in the two‐step procedure as a regressor for bias correction. It has useful derivatives:
\frac{d}{dx}\Bigl[\text{IMR}(x)\Bigr] = \frac{d}{dx}\Bigl[\frac{\phi(x)}{\Phi(x)}\Bigr] = -x\,\text{IMR}(x)\;-\;\bigl[\text{IMR}(x)\bigr]^2.
This arises from the quotient rule and the fact that \phi'(x)=-x\phi(x), \Phi'(x)=\phi(x). The derivative property also helps in interpreting marginal effects in selection models.
36.2.1.2 Treatment Effect (Switching) Model
While the sample selection model is used when outcome is only observed for one group (e.g., D = 1), the treatment effect model is used when outcomes are observed for both groups, but treatment assignment is endogenous.
Treatment Effect Model Equations:
- Outcome: y_i = x_i' \beta + D_i \delta + \varepsilon_i
- Selection: D_i^* = z_i' \gamma + u_i \\ D_i = 1 \text{ if } D_i^* > 0
Where:
D_i is the treatment indicator.
(\varepsilon_i, u_i) are again bivariate normal with correlation \rho.
The treatment effect model is sometimes called a switching regression.
36.2.1.3 Heckman-Type vs. Control Function
- Heckman Sample Selection: Insert an Inverse Mills Ratio (IMR) to adjust the outcome equation for non-random truncation.
- Control Function: Residual-based or predicted-endogenous-variable approach that mirrors IV logic, but typically still requires an instrument or parametric assumption.
| Heckman Sample Selection Model | Heckman-Type Corrections | |
| When | Only observes one sample (treated), addressing selection bias directly. | Two samples are observed (treated and untreated), known as the control function approach. |
| Model | Probit | OLS (even for dummy endogenous variable) |
| Integration of 1st stage | Also include a term (called Inverse Mills ratio) besides the endogenous variable. | Decompose the endogenous variable to get the part that is uncorrelated with the error terms of the outcome equation. Either use the predicted endogenous variable directly or include the residual from the first-stage equation. |
| Advantages and Assumptions | Provides a direct test for endogeneity via the coefficient of the inverse Mills ratio but requires the assumption of joint normality of errors. | Does not require the assumption of joint normality, but can’t test for endogeneity directly. |
36.2.2 Estimation Methods
36.2.2.1 Heckman’s Two-Step Estimator (Heckit)
Step 1: Estimate Selection Equation with Probit
We estimate the probability of being included in the sample: P(w_i = 1 \mid z_i) = \Phi(z_i' \gamma)
From the estimated model, we compute the Inverse Mills Ratio (IMR): \lambda_i = \frac{\phi(z_i' \hat{\gamma})}{\Phi(z_i' \hat{\gamma})}
This term captures the expected value of the error in the outcome equation, conditional on selection.
Step 2: Include IMR in Outcome Equation
We then estimate the regression: y_i = x_i' \beta + \delta \lambda_i + \nu_i
- If \delta is significantly different from 0, selection bias is present.
- \lambda_i corrects for the non-random selection.
- OLS on this augmented model yields consistent estimates of \beta under the joint normality assumption.
- Pros: Conceptually simple; widely used.
- Cons: Relies heavily on the bivariate normal assumption for (\varepsilon_i, u_i). If no good exclusion variable is available, identification rests on the functional form.
Specifically, the model can be identified without an exclusion restriction, but in such cases, identification is driven purely by the non-linearity of the probit function and the normality assumption (through the IMR). This is fragile.
- With strong exclusion restriction for the covariate in the correction equation, the variation in this variable can help identify the control for selection.
- With weak exclusion restriction, and the variable exists in both steps, it’s the assumed error structure that identifies the control for selection (J. Heckman and Navarro-Lozano 2004).
- In management, Wolfolds and Siegel (2019) found that papers should have valid exclusion conditions, because without these, simulations show that results using the Heckman method are less reliable than those obtained with OLS.
For robust identification, we prefer an exclusion restriction:
- A variable that affects selection (through z_i) but not the outcome.
- Example: Distance to a training center might affect the probability of enrollment, but not post-training income.
Without such a variable, the model relies solely on functional form.
The Heckman two-step estimation procedure is less efficient than FIML. One key limitation is that the two-step estimator does not fully exploit the joint distribution of the error terms across equations, leading to a loss of efficiency. Moreover, the two-step approach may introduce measurement error in the second stage. This arises because the inverse Mills ratio used in the second stage is itself an estimated regressor, which can lead to biased standard errors and inference.
###########################
# SIM 1: Heckman 2-step #
###########################
suppressPackageStartupMessages(library(MASS))
set.seed(123)
n <- 1000
rho <- 0.5
beta_true <- 2
gamma_true <- 1.0
Sigma <- matrix(c(1, rho, rho, 1), 2)
errors <- mvrnorm(n, c(0,0), Sigma)
epsilon <- errors[,1]
u <- errors[,2]
x <- rnorm(n)
w <- rnorm(n)
# Selection
z_star <- w*gamma_true + u
z <- ifelse(z_star>0,1,0)
# Outcome
y_star <- x * beta_true + epsilon
# Observed only if z=1
y_obs <- ifelse(z == 1, y_star, NA)
# Step 1: Probit
sel_mod <- glm(z ~ w, family = binomial(link = "probit"))
z_hat <- predict(sel_mod, type = "link")
lambda_vals <- dnorm(z_hat) / pnorm(z_hat)
# Step 2: OLS on observed + IMR
data_heck <- data.frame(y = y_obs,
x = x,
imr = lambda_vals,
z = z)
observed_data <- subset(data_heck, z == 1)
heck_lm <- lm(y ~ x + imr, data = observed_data)
summary(heck_lm)
#>
#> Call:
#> lm(formula = y ~ x + imr, data = observed_data)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.76657 -0.60099 -0.02776 0.56317 2.74797
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.01715 0.07068 0.243 0.808
#> x 1.95925 0.03934 49.800 < 2e-16 ***
#> imr 0.41900 0.10063 4.164 3.69e-05 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 0.8942 on 501 degrees of freedom
#> Multiple R-squared: 0.8332, Adjusted R-squared: 0.8325
#> F-statistic: 1251 on 2 and 501 DF, p-value: < 2.2e-16
cat("True beta=", beta_true, "\n")
#> True beta= 2
cat("Heckman 2-step estimated beta=", coef(heck_lm)["x"], "\n")
#> Heckman 2-step estimated beta= 1.95924936.2.2.2 Full Information Maximum Likelihood
Jointly estimates the selection and outcome equations via ML, assuming:
\biggl(\varepsilon_i, u_i\biggr) \sim \mathcal{N}\biggl(\begin{pmatrix}0\\0\end{pmatrix},\begin{pmatrix}\sigma_{\varepsilon}^2 & \rho\,\sigma_{\varepsilon}\\\rho\,\sigma_{\varepsilon} & 1\end{pmatrix}\biggr).
- Pros: More efficient if the distributional assumption is correct. Allows a direct test of \rho=0 (LR test).
- Cons: More sensitive to specification errors (i.e., requires stronger distributional assumptions); potentially complex to implement.
We can use the sampleSelection package in R to perform full maximum likelihood estimation for the same data:
#############################
# SIM 2: 2-step vs. FIML #
#############################
suppressPackageStartupMessages(library(sampleSelection))
# Using same data (z, x, y_obs) from above
# 1) Heckman 2-step (built-in)
heck2 <-
selection(z ~ w, y_obs ~ x, method = "2step", data = data_heck)
summary(heck2)
#> --------------------------------------------
#> Tobit 2 model (sample selection model)
#> 2-step Heckman / heckit estimation
#> 1000 observations (496 censored and 504 observed)
#> 7 free parameters (df = 994)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.02053 0.04494 0.457 0.648
#> w 0.94063 0.05911 15.913 <2e-16 ***
#> Outcome equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.01715 0.07289 0.235 0.814
#> x 1.95925 0.03924 49.932 <2e-16 ***
#> Multiple R-Squared:0.8332, Adjusted R-Squared:0.8325
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> invMillsRatio 0.4190 0.1018 4.116 4.18e-05 ***
#> sigma 0.9388 NA NA NA
#> rho 0.4463 NA NA NA
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------
# 2) FIML
heckFIML <-
selection(z ~ w, y_obs ~ x, method = "ml", data = data_heck)
summary(heckFIML)
#> --------------------------------------------
#> Tobit 2 model (sample selection model)
#> Maximum Likelihood estimation
#> Newton-Raphson maximisation, 2 iterations
#> Return code 8: successive function values within relative tolerance limit (reltol)
#> Log-Likelihood: -1174.233
#> 1000 observations (496 censored and 504 observed)
#> 6 free parameters (df = 994)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.02169 0.04488 0.483 0.629
#> w 0.94203 0.05908 15.945 <2e-16 ***
#> Outcome equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.008601 0.071315 0.121 0.904
#> x 1.959195 0.039124 50.077 <2e-16 ***
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> sigma 0.94118 0.03503 26.867 < 2e-16 ***
#> rho 0.46051 0.09411 4.893 1.16e-06 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------You can compare the coefficient estimates on x from heck2 vs. heckFIML. In large samples, both should converge to the true \beta. FIML is typically more efficient, but if the normality assumption is violated, both can be biased.
36.2.2.3 CF and IV Approaches
- Control Function
- Residual-based approach: Regress the selection (or treatment) variable on excluded instruments and included controls. Obtain the predicted residual. Include that residual in the main outcome regression.
- If correlated residual is significant, that indicates endogeneity; adjusting for it can correct bias.
- Often used in the context of treatment effect models or simultaneously with IV logic.
- Instrumental Variables
- In the pure treatment effect context, an IV must affect treatment assignment but not the outcome directly.
- For sample selection, an exclusion restriction (“instrument”) must shift selection but not outcomes.
- Example: Distance to a training center influences participation in a job program but not post-training earnings.
36.2.3 Theoretical Connections
36.2.3.1 Conditional Expectation from Truncated Distributions
In the sample selection scenario:
\begin{aligned} \mathbb{E}[y_i\mid w_i=1] &= x_i\beta + \mathbb{E}[\varepsilon_i\mid w_i^*>0],\\ &= x_i\beta + \rho\,\sigma_{\varepsilon}\,\frac{\phi(z_i'\gamma)}{\Phi(z_i'\gamma)}, \end{aligned}
where \rho\,\sigma_{\varepsilon} is the covariance term and \phi and \Phi are the standard normal PDF and CDF, respectively. This formula underpins the inverse Mills ratio correction.
- If \rho>0, then the same unobservables that increase the likelihood of selection also increase outcomes, implying positive selection.
- If \rho<0, selection is negatively correlated with outcomes.
- \hat{\rho}: Estimated correlation of error terms. If significantly different from 0, endogenous selection is present.
- Wald or Likelihood Ratio Test: Used to test H_0: \rho = 0.
- Lambda (\hat{\lambda}): Product of \hat{\rho} \hat{\sigma}_\varepsilon—measures strength of selection bias.
- Inverse Mills Ratio: Can be saved and inspected to understand sample inclusion probabilities.
36.2.3.2 Relationship Among Models
All the models in Unifying Model Frameworks can be seen as special or generalized cases:
If one only has data for the selected group, it’s a sample selection setup.
If data for both groups exist, but assignment is endogenous, it’s a treatment effect problem.
If there’s a valid instrument, one can do a control function or IV approach.
If the normality assumption holds and selection is truly parametric, Heckman or FIML correct for the bias.
Summary Table of Methods
| Method | Data Observed | Key Assumption | Exclusion? | Pros | Cons |
|---|---|---|---|---|---|
| OLS (Naive) | Full or partial, ignoring selection | No endogeneity in errors | Not required | Simple to implement | Biased if endogeneity is present |
| Heckman 2-Step (Heckit) | Outcome only for selected group | Joint normal errors; linear functional | Strongly recommended | Intuitive, widely used | Sensitive to normality/functional form. |
| FIML (Full ML) | Same as Heckman (subset observed) | Joint normal errors | Strongly recommended | More efficient, direct test of \rho=0 | Complex, more sensitive to misspecification |
| Control Function | Observed data for both or one group (depending on setup) | Some form of valid instrument or exog. | Yes (instrument) | Extends easily to many models | Must find valid instrument, no direct test for endogeneity |
| Instrumental Variables | Observed data for both groups, or entire sample (for selection) | IV must affect selection but not outcome | Yes (instrument) | Common approach in program evaluation | Exclusion restriction validity is critical |
36.2.3.3 Concerns
- Small Samples: Two-step procedures can be unstable in smaller datasets.
- Exclusion Restrictions: Without a credible variable that predicts selection but not outcomes, identification depends purely on functional form (bivariate normal + nonlinearity of probit).
- Distributional Assumption: If normality is seriously violated, neither 2-step nor FIML may reliably remove bias.
- Measurement Error in IMR: The second-stage includes an estimated regressor \hat{\lambda}_i, which can add noise.
- Connection to IV: If a strong instrument exists, one could proceed with a control function or standard IV in a treatment effect setup. But for sample selection (lack of data on unselected), the Heckman approach is more common.
- Presence of Correlation between the Error Terms: The Heckman treatment effect model outperforms OLS when \rho \neq 0 because it corrects for selection bias due to unobserved factors. However, when \rho = 0, the correction is unnecessary and can introduce inefficiency, making simpler methods more accurate.
36.2.4 Tobit-2: Heckman’s Sample Selection Model
The Tobit-2 model, also known as Heckman’s standard sample selection model, is designed to correct for sample selection bias. This arises when the outcome variable is only observed for a non-random subset of the population, and the selection process is correlated with the outcome of interest.
A key assumption of the model is the joint normality of the error terms in the selection and outcome equations.
36.2.4.1 Panel Study of Income Dynamics
We demonstrate the model using the classic dataset from Mroz (1984), which provides data from the 1975 Panel Study of Income Dynamics on married women’s labor-force participation and wages.
We aim to estimate the log of hourly wages for married women, using:
-
educ: Years of education -
exper: Years of work experience -
exper^2: Experience squared (to capture non-linear effects) -
city: A dummy for residence in a big city
However, wages are only observed for those who participated in the labor force, meaning an OLS regression using only this subsample would suffer from selection bias.
Because we also have data on non-participants, we can use Heckman’s two-step method to correct for this bias.
- Load and Prepare Data
library(sampleSelection)
library(dplyr)
library(nnet)
library(ggplot2)
library(reshape2)
data("Mroz87") # PSID data on married women in 1975
Mroz87 = Mroz87 %>%
mutate(kids = kids5 + kids618) # total number of children
head(Mroz87)
#> lfp hours kids5 kids618 age educ wage repwage hushrs husage huseduc huswage
#> 1 1 1610 1 0 32 12 3.3540 2.65 2708 34 12 4.0288
#> 2 1 1656 0 2 30 12 1.3889 2.65 2310 30 9 8.4416
#> 3 1 1980 1 3 35 12 4.5455 4.04 3072 40 12 3.5807
#> 4 1 456 0 3 34 12 1.0965 3.25 1920 53 10 3.5417
#> 5 1 1568 1 2 31 14 4.5918 3.60 2000 32 12 10.0000
#> 6 1 2032 0 0 54 12 4.7421 4.70 1040 57 11 6.7106
#> faminc mtr motheduc fatheduc unem city exper nwifeinc wifecoll huscoll
#> 1 16310 0.7215 12 7 5.0 0 14 10.910060 FALSE FALSE
#> 2 21800 0.6615 7 7 11.0 1 5 19.499981 FALSE FALSE
#> 3 21040 0.6915 12 7 5.0 0 15 12.039910 FALSE FALSE
#> 4 7300 0.7815 7 7 5.0 0 6 6.799996 FALSE FALSE
#> 5 27300 0.6215 12 14 9.5 1 7 20.100058 TRUE FALSE
#> 6 19495 0.6915 14 7 7.5 1 33 9.859054 FALSE FALSE
#> kids
#> 1 1
#> 2 2
#> 3 4
#> 4 3
#> 5 3
#> 6 0- Model Overview
The two-step Heckman selection model proceeds as follows:
Selection Equation (Probit):
Models the probability of labor force participation (lfp = 1) as a function of variables that affect the decision to work.Outcome Equation (Wage):
Models log wages conditional on working. A correction term, the IMR, is included to account for the non-random selection into work.
Step 1: Naive OLS on Observed Wages
ols1 = lm(log(wage) ~ educ + exper + I(exper ^ 2) + city,
data = subset(Mroz87, lfp == 1))
summary(ols1)
#>
#> Call:
#> lm(formula = log(wage) ~ educ + exper + I(exper^2) + city, data = subset(Mroz87,
#> lfp == 1))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -3.10084 -0.32453 0.05292 0.36261 2.34806
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.5308476 0.1990253 -2.667 0.00794 **
#> educ 0.1057097 0.0143280 7.378 8.58e-13 ***
#> exper 0.0410584 0.0131963 3.111 0.00199 **
#> I(exper^2) -0.0007973 0.0003938 -2.025 0.04352 *
#> city 0.0542225 0.0680903 0.796 0.42629
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 0.6667 on 423 degrees of freedom
#> Multiple R-squared: 0.1581, Adjusted R-squared: 0.1501
#> F-statistic: 19.86 on 4 and 423 DF, p-value: 5.389e-15This OLS is biased because it only includes women who chose to work.
Step 2: Heckman Two-Step Estimation
# Heckman 2-step estimation
heck1 = heckit(
selection = lfp ~ age + I(age ^ 2) + kids + huswage + educ,
outcome = log(wage) ~ educ + exper + I(exper ^ 2) + city,
data = Mroz87
)
# Stage 1: Selection equation (probit)
summary(heck1$probit)
#> --------------------------------------------
#> Probit binary choice model/Maximum Likelihood estimation
#> Newton-Raphson maximisation, 4 iterations
#> Return code 1: gradient close to zero (gradtol)
#> Log-Likelihood: -482.8212
#> Model: Y == '1' in contrary to '0'
#> 753 observations (325 'negative' and 428 'positive') and 6 free parameters (df = 747)
#> Estimates:
#> Estimate Std. error t value Pr(> t)
#> XS(Intercept) -4.18146681 1.40241567 -2.9816 0.002867 **
#> XSage 0.18608901 0.06517476 2.8552 0.004301 **
#> XSI(age^2) -0.00241491 0.00075857 -3.1835 0.001455 **
#> XSkids -0.14955977 0.03825079 -3.9100 9.230e-05 ***
#> XShuswage -0.04303635 0.01220791 -3.5253 0.000423 ***
#> XSeduc 0.12502818 0.02277645 5.4894 4.034e-08 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> Significance test:
#> chi2(5) = 64.10407 (p=1.719042e-12)
#> --------------------------------------------
# Stage 2: Wage equation with selection correction
summary(heck1$lm)
#>
#> Call:
#> lm(formula = YO ~ -1 + XO + imrData$IMR1, subset = YS == 1, weights = weightsNoNA)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -3.09494 -0.30953 0.05341 0.36530 2.34770
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> XO(Intercept) -0.6143381 0.3768796 -1.630 0.10383
#> XOeduc 0.1092363 0.0197062 5.543 5.24e-08 ***
#> XOexper 0.0419205 0.0136176 3.078 0.00222 **
#> XOI(exper^2) -0.0008226 0.0004059 -2.026 0.04335 *
#> XOcity 0.0510492 0.0692414 0.737 0.46137
#> imrData$IMR1 0.0551177 0.2111916 0.261 0.79423
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 0.6674 on 422 degrees of freedom
#> Multiple R-squared: 0.7734, Adjusted R-squared: 0.7702
#> F-statistic: 240 on 6 and 422 DF, p-value: < 2.2e-16The variable kids is used only in the selection equation. This follows good practice: at least one variable should appear only in the selection equation (serving as an instrument) to help identify the model.
# ML estimation of Heckman selection model
ml1 = selection(
selection = lfp ~ age + I(age^2) + kids + huswage + educ,
outcome = log(wage) ~ educ + exper + I(exper^2) + city,
data = Mroz87
)
summary(ml1)
#> --------------------------------------------
#> Tobit 2 model (sample selection model)
#> Maximum Likelihood estimation
#> Newton-Raphson maximisation, 3 iterations
#> Return code 8: successive function values within relative tolerance limit (reltol)
#> Log-Likelihood: -914.0777
#> 753 observations (325 censored and 428 observed)
#> 13 free parameters (df = 740)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -4.1484037 1.4109302 -2.940 0.003382 **
#> age 0.1842132 0.0658041 2.799 0.005253 **
#> I(age^2) -0.0023925 0.0007664 -3.122 0.001868 **
#> kids -0.1488158 0.0384888 -3.866 0.000120 ***
#> huswage -0.0434253 0.0123229 -3.524 0.000451 ***
#> educ 0.1255639 0.0229229 5.478 5.91e-08 ***
#> Outcome equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.5814781 0.3052031 -1.905 0.05714 .
#> educ 0.1078481 0.0172998 6.234 7.63e-10 ***
#> exper 0.0415752 0.0133269 3.120 0.00188 **
#> I(exper^2) -0.0008125 0.0003974 -2.044 0.04129 *
#> city 0.0522990 0.0682652 0.766 0.44385
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> sigma 0.66326 0.02309 28.729 <2e-16 ***
#> rho 0.05048 0.23169 0.218 0.828
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------The MLE approach jointly estimates both equations, yielding consistent and asymptotically efficient estimates.
Manual Implementation: Constructing the IMR
# Step 1: Probit model
myprob <- probit(lfp ~ age + I(age^2) + kids + huswage + educ,
data = Mroz87)
summary(myprob)
#> --------------------------------------------
#> Probit binary choice model/Maximum Likelihood estimation
#> Newton-Raphson maximisation, 4 iterations
#> Return code 1: gradient close to zero (gradtol)
#> Log-Likelihood: -482.8212
#> Model: Y == '1' in contrary to '0'
#> 753 observations (325 'negative' and 428 'positive') and 6 free parameters (df = 747)
#> Estimates:
#> Estimate Std. error t value Pr(> t)
#> (Intercept) -4.18146681 1.40241567 -2.9816 0.002867 **
#> age 0.18608901 0.06517476 2.8552 0.004301 **
#> I(age^2) -0.00241491 0.00075857 -3.1835 0.001455 **
#> kids -0.14955977 0.03825079 -3.9100 9.230e-05 ***
#> huswage -0.04303635 0.01220791 -3.5253 0.000423 ***
#> educ 0.12502818 0.02277645 5.4894 4.034e-08 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> Significance test:
#> chi2(5) = 64.10407 (p=1.719042e-12)
#> --------------------------------------------
# Step 2: Compute IMR
imr <- invMillsRatio(myprob)
Mroz87$IMR1 <- imr$IMR1
# Step 3: Wage regression including IMR
manually_est <- lm(log(wage) ~ educ + exper + I(exper^2) + city + IMR1,
data = Mroz87,
subset = (lfp == 1))
summary(manually_est)
#>
#> Call:
#> lm(formula = log(wage) ~ educ + exper + I(exper^2) + city + IMR1,
#> data = Mroz87, subset = (lfp == 1))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -3.09494 -0.30953 0.05341 0.36530 2.34770
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.6143381 0.3768796 -1.630 0.10383
#> educ 0.1092363 0.0197062 5.543 5.24e-08 ***
#> exper 0.0419205 0.0136176 3.078 0.00222 **
#> I(exper^2) -0.0008226 0.0004059 -2.026 0.04335 *
#> city 0.0510492 0.0692414 0.737 0.46137
#> IMR1 0.0551177 0.2111916 0.261 0.79423
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 0.6674 on 422 degrees of freedom
#> Multiple R-squared: 0.1582, Adjusted R-squared: 0.1482
#> F-statistic: 15.86 on 5 and 422 DF, p-value: 2.505e-14Equivalent Method Using glm and Manual IMR Calculation
# Probit via glm
probit_selection <- glm(
lfp ~ age + I(age^2) + kids + huswage + educ,
data = Mroz87,
family = binomial(link = 'probit')
)
# Compute predicted latent index and IMR
probit_lp <- -predict(probit_selection)
inv_mills <- dnorm(probit_lp) / (1 - pnorm(probit_lp))
Mroz87$inv_mills <- inv_mills
# Second stage: Wage regression with correction
probit_outcome <- glm(
log(wage) ~ educ + exper + I(exper^2) + city + inv_mills,
data = Mroz87,
subset = (lfp == 1)
)
summary(probit_outcome)
#>
#> Call:
#> glm(formula = log(wage) ~ educ + exper + I(exper^2) + city +
#> inv_mills, data = Mroz87, subset = (lfp == 1))
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.6143383 0.3768798 -1.630 0.10383
#> educ 0.1092363 0.0197062 5.543 5.24e-08 ***
#> exper 0.0419205 0.0136176 3.078 0.00222 **
#> I(exper^2) -0.0008226 0.0004059 -2.026 0.04335 *
#> city 0.0510492 0.0692414 0.737 0.46137
#> inv_mills 0.0551179 0.2111918 0.261 0.79423
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> (Dispersion parameter for gaussian family taken to be 0.4454809)
#>
#> Null deviance: 223.33 on 427 degrees of freedom
#> Residual deviance: 187.99 on 422 degrees of freedom
#> AIC: 876.49
#>
#> Number of Fisher Scoring iterations: 2Comparing Models
library(stargazer)
library(plm)
library(sandwich)
# Custom robust SE function
cse = function(reg) {
sqrt(diag(vcovHC(reg, type = "HC1")))
}
# Comparison table
stargazer(
ols1, heck1, ml1, manually_est,
se = list(cse(ols1), NULL, NULL, NULL),
title = "Married Women's Wage Regressions: OLS vs Heckman Models",
type = "text",
df = FALSE,
digits = 4,
selection.equation = TRUE
)
#>
#> Married Women's Wage Regressions: OLS vs Heckman Models
#> =========================================================================
#> Dependent variable:
#> -----------------------------------------------------
#> log(wage) lfp log(wage)
#> OLS Heckman selection OLS
#> selection
#> (1) (2) (3) (4)
#> -------------------------------------------------------------------------
#> age 0.1861*** 0.1842***
#> (0.0652) (0.0658)
#>
#> I(age2) -0.0024*** -0.0024***
#> (0.0008) (0.0008)
#>
#> kids -0.1496*** -0.1488***
#> (0.0383) (0.0385)
#>
#> huswage -0.0430*** -0.0434***
#> (0.0122) (0.0123)
#>
#> educ 0.1057*** 0.1250*** 0.1256*** 0.1092***
#> (0.0130) (0.0228) (0.0229) (0.0197)
#>
#> exper 0.0411*** 0.0419***
#> (0.0154) (0.0136)
#>
#> I(exper2) -0.0008* -0.0008**
#> (0.0004) (0.0004)
#>
#> city 0.0542 0.0510
#> (0.0653) (0.0692)
#>
#> IMR1 0.0551
#> (0.2112)
#>
#> Constant -0.5308*** -4.1815*** -4.1484*** -0.6143
#> (0.2032) (1.4024) (1.4109) (0.3769)
#>
#> -------------------------------------------------------------------------
#> Observations 428 753 753 428
#> R2 0.1581 0.1582 0.1582
#> Adjusted R2 0.1501 0.1482 0.1482
#> Log Likelihood -914.0777
#> rho 0.0830 0.0505 (0.2317)
#> Inverse Mills Ratio 0.0551 (0.2099)
#> Residual Std. Error 0.6667 0.6674
#> F Statistic 19.8561*** 15.8635***
#> =========================================================================
#> Note: *p<0.1; **p<0.05; ***p<0.01IMR: If the coefficient on the IMR is statistically significant, it suggests selection bias is present and that OLS estimates are biased.
\rho: Represents the estimated correlation between the error terms of the selection and outcome equations. A significant \rho implies non-random selection, justifying the Heckman correction.
In our case, if the IMR coefficient is not statistically different from zero, then selection bias may not be a serious concern.
36.2.4.2 The Role of Exclusion Restrictions in Heckman’s Model
This example, adapted from the sampleSelection, demonstrates the identification of the sample selection model using simulated data.
We compare two cases:
One where an exclusion restriction is present (i.e., selection and outcome equations use different regressors).
One where the same regressor is used in both equations.
Case 1: With Exclusion Restriction
set.seed(0)
library(sampleSelection)
library(mvtnorm)
# Simulate bivariate normal error terms with correlation -0.7
eps <-
rmvnorm(500, mean = c(0, 0), sigma = matrix(c(1,-0.7,-0.7, 1), 2, 2))
# Independent explanatory variable for selection equation
xs <- runif(500)
# Selection: Probit model (latent utility model)
ys_latent <- xs + eps[, 1]
ys <- ys_latent > 0 # observed participation indicator (TRUE/FALSE)
# Independent explanatory variable for outcome equation
xo <- runif(500)
# Latent outcome variable
yo_latent <- xo + eps[, 2]
# Observed outcome: only when selected (ys == TRUE)
yo <- yo_latent * ys
# Estimate Heckman's selection model
model_with_exclusion <-
selection(selection = ys ~ xs, outcome = yo ~ xo)
summary(model_with_exclusion)
#> --------------------------------------------
#> Tobit 2 model (sample selection model)
#> Maximum Likelihood estimation
#> Newton-Raphson maximisation, 5 iterations
#> Return code 1: gradient close to zero (gradtol)
#> Log-Likelihood: -712.3163
#> 500 observations (172 censored and 328 observed)
#> 6 free parameters (df = 494)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.2228 0.1081 -2.061 0.0399 *
#> xs 1.3377 0.2014 6.642 8.18e-11 ***
#> Outcome equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.0002265 0.1294178 -0.002 0.999
#> xo 0.7299070 0.1635925 4.462 1.01e-05 ***
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> sigma 0.9190 0.0574 16.009 < 2e-16 ***
#> rho -0.5392 0.1521 -3.544 0.000431 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------Key Observations:
The variables
xsandxoare independent, fulfilling the exclusion restriction.The outcome equation is identified not only by the non-linearity of the model but also by the presence of a regressor in the selection equation that is absent from the outcome equation.
This mirrors realistic scenarios in applied business settings, such as:
Participation in the labor force driven by family or geographic factors (
xs), while wages depend on skills or education (xo).Loan application driven by personal risk preferences, while interest rates depend on credit score or income.
Case 2: Without Exclusion Restriction
# Now use the same regressor (xs) in both equations
yo_latent2 <- xs + eps[, 2]
yo2 <- yo_latent2 * ys
# Re-estimate model without exclusion restriction
model_no_exclusion <-
selection(selection = ys ~ xs, outcome = yo2 ~ xs)
summary(model_no_exclusion)
#> --------------------------------------------
#> Tobit 2 model (sample selection model)
#> Maximum Likelihood estimation
#> Newton-Raphson maximisation, 14 iterations
#> Return code 8: successive function values within relative tolerance limit (reltol)
#> Log-Likelihood: -712.8298
#> 500 observations (172 censored and 328 observed)
#> 6 free parameters (df = 494)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.1984 0.1114 -1.781 0.0756 .
#> xs 1.2907 0.2085 6.191 1.25e-09 ***
#> Outcome equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.5499 0.5644 -0.974 0.33038
#> xs 1.3987 0.4482 3.120 0.00191 **
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> sigma 0.85091 0.05352 15.899 <2e-16 ***
#> rho -0.13226 0.72684 -0.182 0.856
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------What changes?
The estimates remain approximately unbiased because the model is still identified by functional form (non-linear selection mechanism).
However, the standard errors are substantially larger, leading to less precise inference.
Why Exclusion Restrictions Matter
The exclusion restriction improves identification by introducing additional variation that affects selection but not the outcome. This is a common recommendation in empirical work:
In the first case,
xshelps explain who is selected, whilexohelps explain the outcome, enabling more precise estimates.In the second case, all variation comes from
xs, meaning the model relies solely on the distributional assumptions (e.g., joint normality of errors) for identification.
Best Practice (for applied researchers):
Always try to include at least one variable in the selection equation that does not appear in the outcome equation. This helps identify the model and improves estimation precision.
36.2.5 Tobit-5: Switching Regression Model
The Tobit-5 model, also known as the Switching Regression Model, generalizes Heckman’s sample selection model to allow for two separate outcome equations:
- One model for participants
- A different model for non-participants
Assumptions:
- The selection process determines which outcome equation is observed.
- There is at least one variable in the selection equation that does not appear in the outcome equations (i.e., an exclusion restriction), which improves identification.
- The errors from all three equations (selection, outcome1, outcome2) are assumed to be jointly normally distributed.
This model is especially relevant when selection is endogenous and both groups (selected and unselected) have distinct data-generating processes.
36.2.5.1 Simulated Example: With Exclusion Restriction
set.seed(0)
library(sampleSelection)
library(mvtnorm)
# Define a 3x3 covariance matrix with positive correlations
vc <- diag(3)
vc[lower.tri(vc)] <- c(0.9, 0.5, 0.1)
vc[upper.tri(vc)] <- t(vc)[upper.tri(vc)]
# Generate multivariate normal error terms
eps <- rmvnorm(500, mean = c(0, 0, 0), sigma = vc)
# Selection equation regressor (xs), uniformly distributed
xs <- runif(500)
# Binary selection indicator: ys = 1 if selected
ys <- xs + eps[, 1] > 0
# Separate regressors for two regimes (fulfilling exclusion restriction)
xo1 <- runif(500)
yo1 <- xo1 + eps[, 2] # Outcome for selected group
xo2 <- runif(500)
yo2 <- xo2 + eps[, 3] # Outcome for non-selected group
# Fit switching regression model
model_switch <- selection(ys ~ xs, list(yo1 ~ xo1, yo2 ~ xo2))
summary(model_switch)
#> --------------------------------------------
#> Tobit 5 model (switching regression model)
#> Maximum Likelihood estimation
#> Newton-Raphson maximisation, 11 iterations
#> Return code 1: gradient close to zero (gradtol)
#> Log-Likelihood: -895.8201
#> 500 observations: 172 selection 1 (FALSE) and 328 selection 2 (TRUE)
#> 10 free parameters (df = 490)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.1550 0.1051 -1.474 0.141
#> xs 1.1408 0.1785 6.390 3.86e-10 ***
#> Outcome equation 1:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.02708 0.16395 0.165 0.869
#> xo1 0.83959 0.14968 5.609 3.4e-08 ***
#> Outcome equation 2:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.1583 0.1885 0.840 0.401
#> xo2 0.8375 0.1707 4.908 1.26e-06 ***
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> sigma1 0.93191 0.09211 10.118 <2e-16 ***
#> sigma2 0.90697 0.04434 20.455 <2e-16 ***
#> rho1 0.88988 0.05353 16.623 <2e-16 ***
#> rho2 0.17695 0.33139 0.534 0.594
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------The estimated coefficients are close to the true values (intercept = 0, slope = 1), and the model converges well due to correct specification and valid exclusion restriction.
36.2.5.2 Example: Functional Form Misspecification
To demonstrate how non-normal errors or skewed distributions affect the model, consider the following:
set.seed(0)
eps <- rmvnorm(1000, mean = rep(0, 3), sigma = vc)
# Induce skewness: squared errors minus 1 to ensure mean zero
eps <- eps^2 - 1
# Generate xs on a skewed interval [-1, 0]
xs <- runif(1000, -1, 0)
ys <- xs + eps[, 1] > 0
xo1 <- runif(1000)
yo1 <- xo1 + eps[, 2]
xo2 <- runif(1000)
yo2 <- xo2 + eps[, 3]
# Attempt model estimation
summary(selection(ys ~ xs, list(yo1 ~ xo1, yo2 ~ xo2), iterlim = 20))
#> --------------------------------------------
#> Tobit 5 model (switching regression model)
#> Maximum Likelihood estimation
#> Newton-Raphson maximisation, 12 iterations
#> Return code 3: Last step could not find a value above the current.
#> Boundary of parameter space?
#> Consider switching to a more robust optimisation method temporarily.
#> Log-Likelihood: -1695.102
#> 1000 observations: 782 selection 1 (FALSE) and 218 selection 2 (TRUE)
#> 10 free parameters (df = 990)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.660315 0.082477 -8.006 3.3e-15 ***
#> xs 0.007167 0.088630 0.081 0.936
#> Outcome equation 1:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.31351 0.04868 -6.44 1.86e-10 ***
#> xo1 1.03862 0.08049 12.90 < 2e-16 ***
#> Outcome equation 2:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -2.6835 0.2043 -13.132 < 2e-16 ***
#> xo2 1.0230 0.1309 7.814 1.41e-14 ***
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> sigma1 0.70172 0.02000 35.09 <2e-16 ***
#> sigma2 2.49651 NaN NaN NaN
#> rho1 0.51564 0.04216 12.23 <2e-16 ***
#> rho2 1.00000 NaN NaN NaN
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------Even though the exclusion restriction is preserved, non-normal errors introduce bias in the intercept estimates, and convergence is less reliable. This illustrates how functional form misspecification (i.e., deviations from assumed distributional forms) impacts performance.
36.2.5.3 Example: No Exclusion Restriction
Here we remove the exclusion restriction by using the same regressor (xs) in both outcome equations.
set.seed(0)
xs <- runif(1000, -1, 1)
ys <- xs + eps[, 1] > 0
yo1 <- xs + eps[, 2]
yo2 <- xs + eps[, 3]
# Fit switching regression without exclusion restriction
summary(tmp <-
selection(ys ~ xs, list(yo1 ~ xs, yo2 ~ xs), iterlim = 20))
#> --------------------------------------------
#> Tobit 5 model (switching regression model)
#> Maximum Likelihood estimation
#> Newton-Raphson maximisation, 16 iterations
#> Return code 1: gradient close to zero (gradtol)
#> Log-Likelihood: -1879.552
#> 1000 observations: 615 selection 1 (FALSE) and 385 selection 2 (TRUE)
#> 10 free parameters (df = 990)
#> Probit selection equation:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.33425 0.04280 -7.81 1.46e-14 ***
#> xs 0.94762 0.07763 12.21 < 2e-16 ***
#> Outcome equation 1:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.49592 0.06800 -7.293 6.19e-13 ***
#> xs 0.84530 0.06789 12.450 < 2e-16 ***
#> Outcome equation 2:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.3861 0.4967 0.777 0.4371
#> xs 0.6254 0.3322 1.882 0.0601 .
#> Error terms:
#> Estimate Std. Error t value Pr(>|t|)
#> sigma1 0.61693 0.02054 30.029 <2e-16 ***
#> sigma2 1.59059 0.05745 27.687 <2e-16 ***
#> rho1 0.19981 0.15863 1.260 0.208
#> rho2 -0.01259 0.29339 -0.043 0.966
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#> --------------------------------------------This model may fail to converge or produce biased estimates, especially for the intercepts. Even if it does converge, the reliability of the inference is questionable due to weak identification from using the same regressor across all equations.
Notes on Estimation and Convergence
The log-likelihood function of switching regression models is not globally concave, so the estimation process may:
Fail to converge
Converge to a local maximum
Practical Tips:
Try different starting values or random seeds
Use alternative maximization algorithms (e.g.,
optimcontrol)Consider rescaling variables or centering predictors
Refer to Non-Linear Regression for advanced diagnostics
Model Comparison Summary
| Scenario | Exclusion Restriction | Convergence | Bias | Variance |
|---|---|---|---|---|
| Well-specified with exclusion | Yes | Likely | No | Low |
| Misspecified distribution | Yes | Risky | Yes (intercepts) | Moderate |
| No exclusion restriction | No | Often fails | Yes | High |
36.2.6 Pattern-Mixture Models
In the context of endogenous sample selection, one of the central challenges is modeling the joint distribution of the outcome and the selection mechanism when data are Missing Not At Random (MNAR). In this framework, the probability that an outcome is observed may depend on unobserved values of that outcome, making the missingness mechanism nonignorable.
Previously, we discussed the selection model approach (e.g., Heckman’s Tobit-2 model), which factorizes the joint distribution as:
\mathbb{P}(Y, R) = \mathbb{P}(Y) \cdot \mathbb{P}(R \mid Y),
where:
Y is the outcome of interest,
R is the response indicator (with R = 1 if Y is observed, R = 0 otherwise).
This approach models the selection process explicitly via \mathbb{P}(R \mid Y), often using a parametric model such as the probit.
However, the pattern-mixture model (PMM) offers an alternative and equally valid factorization:
\mathbb{P}(Y, R) = \mathbb{P}(Y \mid R) \cdot \mathbb{P}(R),
which decomposes the joint distribution by conditioning on the response pattern. This approach is particularly advantageous when the selection mechanism is complex, or when interest lies in modeling how the outcome distribution varies across response strata.
36.2.6.1 Definition of the Pattern-Mixture Model
Let (Y_i, R_i) for i = 1, \dots, n denote the observed data, where:
- Y_i \in \mathbb{R}^p is the multivariate outcome of interest,
- R_i \in \{0,1\}^p is a missingness pattern indicator vector, where R_{ij} = 1 indicates that Y_{ij} is observed.
Define \mathcal{R} as the finite set of all possible response patterns. For each pattern r \in \mathcal{R}, partition the outcome vector Y into:
- Y_{(r)}: observed components of Y under pattern r,
- Y_{(\bar{r})}: missing components of Y under pattern r.
The joint distribution can then be factorized according to the pattern-mixture model as:
f(Y, R) = f(Y \mid R = r) \cdot \mathbb{P}(R = r), \quad r \in \mathcal{R}.
The marginal distribution of Y is obtained by summing over all response patterns:
f(Y) = \sum_{r \in \mathcal{R}} f(Y \mid R = r) \cdot \mathbb{P}(R = r).
In the simplest case of binary response patterns (complete responders R=1 and complete nonresponders R=0), this reduces to:
\mathbb{P}(Y) = \mathbb{P}(Y \mid R = 1) \cdot \mathbb{P}(R = 1) + \mathbb{P}(Y \mid R = 0) \cdot \mathbb{P}(R = 0).
Thus, the overall distribution of Y is explicitly treated as a mixture of distributions for responders and nonresponders. While \mathbb{P}(Y \mid R = 1) can be directly estimated from observed data, \mathbb{P}(Y \mid R = 0) cannot be identified from data alone, necessitating additional assumptions or sensitivity parameters for its estimation.
Common approaches for addressing non-identifiability involve defining plausible deviations from \mathbb{P}(Y \mid R = 1) using specific sensitivity parameters such as:
- Shift Bias: Adjusting imputed values by a constant \delta to reflect systematic differences in nonresponders.
- Scale Bias: Scaling imputed values to account for differing variability.
- Shape Bias: Reweighting imputations, for example, using methods like Selection-Indicator Reweighting (SIR), to capture distributional shape differences.
In this discussion, primary focus is placed on the shift parameter \delta, hypothesizing that nonresponse shifts the mean of Y by \delta units.
In practice, since only observed components Y_{(r)} are available for individuals with missing data, modeling the full conditional distribution f(Y \mid R = r) requires extrapolating from observed to unobserved components.
36.2.6.2 Modeling Strategy and Identifiability
Suppose the full-data conditional distribution is parameterized by pattern-specific parameters \theta_r for each response pattern r \in \mathcal{R}:
f(Y \mid R = r; \theta_r), \quad \text{with } \theta = \{\theta_r : r \in \mathcal{R}\}.
Then, the marginal distribution of Y can be expressed as:
f(Y; \theta, \psi) = \sum_{r \in \mathcal{R}} f(Y \mid R = r; \theta_r) \cdot \mathbb{P}(R = r; \psi),
where \psi parameterizes the distribution of response patterns.
Important points about identifiability include:
- The model is inherently non-identifiable without explicit assumptions regarding the distribution of missing components Y_{(\bar{r})}.
- Estimating f(Y_{(\bar{r})} \mid Y_{(r)}, R = r) requires external information, expert knowledge, or assumptions encapsulated in sensitivity parameters.
The conditional density for each response pattern can be further decomposed as:
f(Y \mid R = r) = f(Y_{(r)} \mid R = r) \cdot f(Y_{(\bar{r})} \mid Y_{(r)}, R = r).
- The first term f(Y_{(r)} \mid R = r) is estimable directly from observed data.
- The second term f(Y_{(\bar{r})} \mid Y_{(r)}, R = r) cannot be identified from observed data alone and must rely on assumptions or additional sensitivity analysis.
36.2.6.3 Location Shift Models
A widely used strategy in PMMs is to introduce a location shift for the unobserved outcomes. Specifically, assume that:
Y_{(\bar{r})} \mid Y_{(r)}, R = r \sim \mathcal{L}(Y_{(\bar{r})} \mid Y_{(r)}, R = r = r^*) + \delta_r,
where:
r^* denotes a fully observed (reference) pattern (e.g., R = 1^p),
\delta_r is a vector of sensitivity parameters that quantify the mean shift for missing outcomes under pattern r.
More formally, for each r \in \mathcal{R} and unobserved component j \in \bar{r}, we specify:
\mathbb{E}[Y_j \mid Y_{(r)}, R = r] = \mathbb{E}[Y_j \mid Y_{(r)}, R = r^*] + \delta_{rj}.
Under this framework:
- Setting \delta_r = 0 corresponds to the MAR assumption (i.e., ignorability).
- Nonzero \delta_r introduces controlled deviations from MAR and enables sensitivity analysis.
36.2.6.4 Sensitivity Analysis in Pattern-Mixture Models
To evaluate the robustness of inferences to the missing data mechanism, we perform sensitivity analysis by varying \delta_r over plausible ranges.
Bias in Estimation
Let \mu = \mathbb{E}[Y] be the target estimand. Under the pattern-mixture decomposition:
\mu = \sum_{r \in \mathcal{R}} \mathbb{E}[Y \mid R = r] \cdot \mathbb{P}(R = r),
and under the shift model:
\mathbb{E}[Y \mid R = r] = \tilde{\mu}_r + \delta_r^{*},
where:
\tilde{\mu}_r is the mean computed using observed data (e.g., via imputation under MAR),
\delta_r^{*} is a vector with entries equal to the mean shifts \delta_{rj} for missing components and 0 for observed components.
Therefore, the overall bias induced by nonzero shifts is:
\Delta = \sum_{r \in \mathcal{R}} \delta_r^{*} \cdot \mathbb{P}(R = r).
This highlights how the overall expectation depends linearly on the sensitivity parameters and their associated pattern probabilities.
Practical Recommendations
- Use subject-matter knowledge to specify plausible ranges for \delta_r.
- Consider standard increments (e.g., \pm 0.2 SD units) or domain-specific metrics (e.g., \pm 5 P/E for valuation).
- If the dataset is large and the number of missingness patterns is small (e.g., monotone dropout), more detailed pattern-specific modeling is feasible.
36.2.6.5 Generalization to Multivariate and Longitudinal Data
Suppose Y = (Y_1, \dots, Y_T) is a longitudinal outcome. Let D \in \{1, \dots, T+1\} denote the dropout time, where D = t implies observation up to time t-1.
Then the pattern-mixture factorization becomes:
f(Y, D) = f(Y \mid D = t) \cdot \mathbb{P}(D = t), \quad t = 2, \dots, T+1.
Assume a parametric model:
Y_j \mid D = t \sim \mathcal{N}(\beta_{0t} + \beta_{1t} t_j, \sigma_t^2), \quad \text{for } j = 1, \dots, T.
Then the overall mean trajectory becomes:
\mathbb{E}[Y_j] = \sum_{t=2}^{T+1} (\beta_{0t} + \beta_{1t} t_j) \cdot \mathbb{P}(D = t).
This model allows for dropout-pattern-specific trajectories and can flexibly account for deviations in the distributional shape due to dropout.
36.2.6.6 Comparison with Selection Models
| Feature | Selection Model | Pattern-Mixture Model |
|---|---|---|
| Factorization | \mathbb{P}(Y) \cdot \mathbb{P}(R \mid Y) | \mathbb{P}(Y \mid R) \cdot \mathbb{P}(R) |
| Target of modeling | Missingness process \mathbb{P}(R \mid Y) | Distribution of Y under R |
| Assumption for identifiability | MAR (often Probit) | Extrapolation (e.g., shift \delta) |
| Modeling burden | On \mathbb{P}(R \mid Y) | On f(Y \mid R) |
| Sensitivity analysis | Less interpretable | Easily parameterized (via \delta) |
library(tidyverse)
library(mice)
library(mvtnorm)
library(ggplot2)
library(broom)
library(patchwork)- Simulate Longitudinal Dropout Data
We simulate a 3-time-point longitudinal outcome, where dropout depends on unobserved future outcomes, i.e., MNAR.
set.seed(123)
# Parameters
n <- 100
time <- c(0, 1, 2)
beta <- c(100, 5) # intercept and slope
sigma <- 5
dropout_bias <- -10 # MNAR: lower future Y -> more dropout
# Simulate full data (Y1, Y2, Y3)
Y <- matrix(NA, nrow = n, ncol = 3)
for (i in 1:3) {
Y[, i] <- beta[1] + beta[2] * time[i] + rnorm(n, 0, sigma)
}
# Dropout mechanism: higher chance to drop if Y3 is low
prob_dropout <- plogis(-0.5 + 0.2 * (dropout_bias - Y[, 3]))
drop <- rbinom(n, 1, prob_dropout)
# Define dropout time: if drop == 1, then Y3 is missing
R <- matrix(1, nrow = n, ncol = 3)
R[drop == 1, 3] <- 0
# Observed data
Y_obs <- Y
Y_obs[R == 0] <- NA
# Combine into a data frame
df <-
data.frame(
id = 1:n,
Y1 = Y_obs[, 1],
Y2 = Y_obs[, 2],
Y3 = Y_obs[, 3],
R3 = R[, 3]
)- Fit Pattern-Mixture Model with Delta Adjustment
We model Y_3 as a function of Y_1 and Y_2, using the observed cases (complete cases), and create multiple imputed datasets under various shift parameters \delta to represent different MNAR scenarios.
# Base linear model for Y3 ~ Y1 + Y2
model_cc <- lm(Y3 ~ Y1 + Y2, data = df, subset = R3 == 1)
# Predict for missing Y3s
preds <- predict(model_cc, newdata = df)
# Sensitivity values for delta
delta_seq <- c(0, -2.5, -5, -7.5, -10) # in units of Y3
# Perform delta adjustment
imputed_dfs <- lapply(delta_seq, function(delta) {
df_new <- df
df_new$Y3_imp <- ifelse(is.na(df$Y3),
preds + delta,
df$Y3)
df_new$delta <- delta
return(df_new)
})
# Stack all results
df_imp_all <- bind_rows(imputed_dfs)- Estimate Full-Data Mean and Trajectory under Each Delta
We now estimate the mean of Y_3 and the full outcome trajectory across delta values.
estimates <- df_imp_all %>%
group_by(delta) %>%
summarise(
mu_Y1 = mean(Y1, na.rm = TRUE),
mu_Y2 = mean(Y2, na.rm = TRUE),
mu_Y3 = mean(Y3_imp),
.groups = "drop"
) %>%
pivot_longer(cols = starts_with("mu"),
names_to = "Time",
values_to = "Mean") %>%
mutate(Time = factor(
Time,
levels = c("mu_Y1", "mu_Y2", "mu_Y3"),
labels = c("Time 1", "Time 2", "Time 3")
))- Visualization: Sensitivity Analysis
We visualize how the mean trajectory changes across different sensitivity parameters \delta.
ggplot(estimates, aes(
x = Time,
y = Mean,
color = as.factor(delta),
group = delta
)) +
geom_line(linewidth = 1) +
geom_point(size = 2) +
scale_color_viridis_d(name = expression(delta)) +
labs(
title = "Pattern-Mixture Model Sensitivity Analysis",
y = "Mean of Y_t",
x = "Time"
) +
causalverse::ama_theme()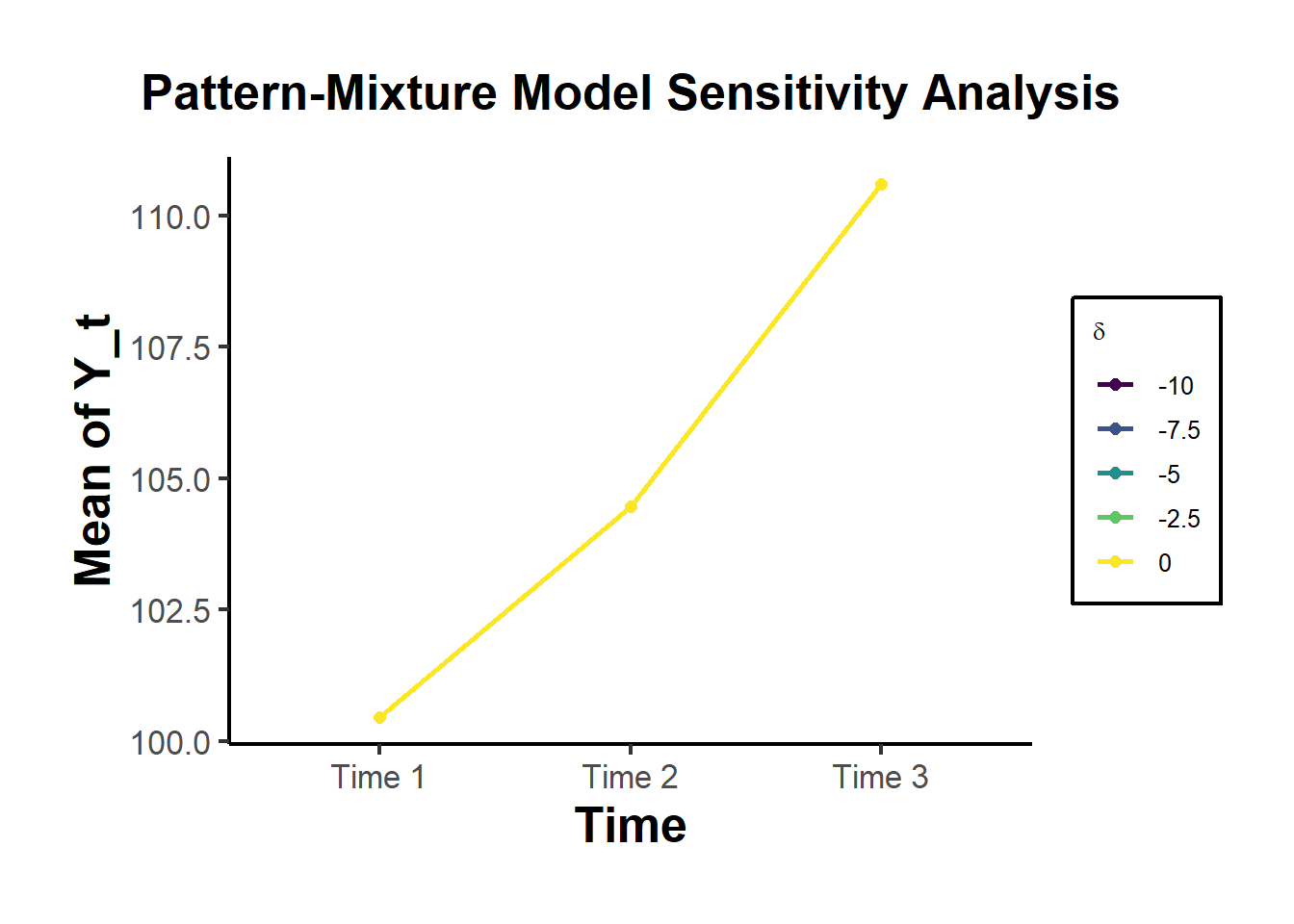
- Sensitivity Table for Reporting
This table quantifies the change in \mu_{Y3} across sensitivity scenarios.
df_summary <- df_imp_all %>%
group_by(delta) %>%
summarise(
Mean_Y3 = mean(Y3_imp),
SD_Y3 = sd(Y3_imp),
N_Missing = sum(is.na(Y3)),
N_Observed = sum(!is.na(Y3))
)
knitr::kable(df_summary, digits = 2,
caption = "Sensitivity of Estimated Mean of Y3 to Delta Adjustments")| delta | Mean_Y3 | SD_Y3 | N_Missing | N_Observed |
|---|---|---|---|---|
| -10.0 | 110.6 | 4.75 | 0 | 100 |
| -7.5 | 110.6 | 4.75 | 0 | 100 |
| -5.0 | 110.6 | 4.75 | 0 | 100 |
| -2.5 | 110.6 | 4.75 | 0 | 100 |
| 0.0 | 110.6 | 4.75 | 0 | 100 |
- Interpretation and Discussion
The estimated mean of Y_3 decreases linearly with increasingly negative \delta, as expected.
This illustrates the non-identifiability of the distribution of missing data without unverifiable assumptions.
The results provide a tipping point analysis: at what \delta does inference about the mean (or treatment effect, in a causal study) substantially change?