4.1 More R basics
In order to implement some of the contents of this chapter we need to cover more R basics, mostly related with flexible plotting that is not implemented directly in R Commander. The R functions we will are also very useful for simplifying some R Commander approaches.
In the following sections, type – not copy and paste systematically – the code in the 'R Script' panel and send it to the output panel. Remember that you should get the same outputs (which are preceded by ## [1]).
4.1.1 Data frames revisited
# Let's begin importing the iris dataset
data(iris)
# names gives you the variables in the data frame
names(iris)
## [1] "Sepal.Length" "Sepal.Width" "Petal.Length" "Petal.Width" "Species"
# The beginning of the data
head(iris)
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 1 5.1 3.5 1.4 0.2 setosa
## 2 4.9 3.0 1.4 0.2 setosa
## 3 4.7 3.2 1.3 0.2 setosa
## 4 4.6 3.1 1.5 0.2 setosa
## 5 5.0 3.6 1.4 0.2 setosa
## 6 5.4 3.9 1.7 0.4 setosa
# So we can access variables by $ or as in a matrix
iris$Sepal.Length[1:10]
## [1] 5.1 4.9 4.7 4.6 5.0 5.4 4.6 5.0 4.4 4.9
iris[1:10, 1]
## [1] 5.1 4.9 4.7 4.6 5.0 5.4 4.6 5.0 4.4 4.9
iris[3, 1]
## [1] 4.7
# Information on the dimension of the data frame
dim(iris)
## [1] 150 5
# str gives the structure of any object in R
str(iris)
## 'data.frame': 150 obs. of 5 variables:
## $ Sepal.Length: num 5.1 4.9 4.7 4.6 5 5.4 4.6 5 4.4 4.9 ...
## $ Sepal.Width : num 3.5 3 3.2 3.1 3.6 3.9 3.4 3.4 2.9 3.1 ...
## $ Petal.Length: num 1.4 1.4 1.3 1.5 1.4 1.7 1.4 1.5 1.4 1.5 ...
## $ Petal.Width : num 0.2 0.2 0.2 0.2 0.2 0.4 0.3 0.2 0.2 0.1 ...
## $ Species : Factor w/ 3 levels "setosa","versicolor",..: 1 1 1 1 1 1 1 1 1 1 ...
# Recall the species variable: it is a categorical variable (or factor),
# not a numeric variable
iris$Species[1:10]
## [1] setosa setosa setosa setosa setosa setosa setosa setosa setosa setosa
## Levels: setosa versicolor virginica
# Factors can only take certain values
levels(iris$Species)
## [1] "setosa" "versicolor" "virginica"
# If a file contains a variable with character strings as observations (either
# encapsulated by quotation marks or not), the variable will become a factor
# when imported into RDo the following:
-
Import
auto.txtinto R as the data frameauto. Check how the character strings in the file give rise to factor variables. -
Get the dimensions of
autoand show beginning of the data. -
Retrieve the fifth observation of
horsepowerin two different ways. -
Compute the levels of
name.
4.1.2 Vector-related functions
# The function seq creates sequences of numbers equally separated
seq(0, 1, by = 0.1)
## [1] 0.0 0.1 0.2 0.3 0.4 0.5 0.6 0.7 0.8 0.9 1.0
seq(0, 1, length.out = 5)
## [1] 0.00 0.25 0.50 0.75 1.00
# You can short the latter argument
seq(0, 1, l = 5)
## [1] 0.00 0.25 0.50 0.75 1.00
# Repeat number
rep(0, 5)
## [1] 0 0 0 0 0
# Reverse a vector
myVec <- c(1:5, -1:3)
rev(myVec)
## [1] 3 2 1 0 -1 5 4 3 2 1
# Another way
myVec[length(myVec):1]
## [1] 3 2 1 0 -1 5 4 3 2 1
# Count repetitions in your data
table(iris$Sepal.Length)
##
## 4.3 4.4 4.5 4.6 4.7 4.8 4.9 5 5.1 5.2 5.3 5.4 5.5 5.6 5.7 5.8 5.9 6 6.1 6.2
## 1 3 1 4 2 5 6 10 9 4 1 6 7 6 8 7 3 6 6 4
## 6.3 6.4 6.5 6.6 6.7 6.8 6.9 7 7.1 7.2 7.3 7.4 7.6 7.7 7.9
## 9 7 5 2 8 3 4 1 1 3 1 1 1 4 1
table(iris$Species)
##
## setosa versicolor virginica
## 50 50 50Do the following:
- Create the vector \(x=(0.3, 0.6, 0.9, 1.2)\).
- Create a vector of length 100 ranging from \(0\) to \(1\) with entries equally separated.
-
Compute the amount of zeros and ones in
x <- c(0, 0, 1, 0, 1, 0, 0, 1, 0, 1, 0). Check that they are the same as inrev(x). -
Compute the vector \((0.1, 1.1, 2.1, ..., 100.1)\) in four different ways using
seqandrev. Do the same but using:instead ofseq. (Hint: add0.1)
4.1.3 Logical conditions and subsetting
# Relational operators: x < y, x > y, x <= y, x >= y, x == y, x!= y
# They return TRUE or FALSE
# Smaller than
0 < 1
## [1] TRUE
# Greater than
1 > 1
## [1] FALSE
# Greater or equal to
1 >= 1 # Remember: ">="" and not "=>"" !
## [1] TRUE
# Smaller or equal to
2 <= 1 # Remember: "<="" and not "=<"" !
## [1] FALSE
# Equal
1 == 1 # Tests equality. Remember: "=="" and not "="" !
## [1] TRUE
# Unequal
1 != 0 # Tests inequality
## [1] TRUE
# TRUE is encoded as 1 and FALSE as 0
TRUE + 1
## [1] 2
FALSE + 1
## [1] 1
# In a vector-like fashion
x <- 1:5
y <- c(0, 3, 1, 5, 2)
x < y
## [1] FALSE TRUE FALSE TRUE FALSE
x == y
## [1] FALSE FALSE FALSE FALSE FALSE
x != y
## [1] TRUE TRUE TRUE TRUE TRUE
# Subsetting of vectors
x
## [1] 1 2 3 4 5
x[x >= 2]
## [1] 2 3 4 5
x[x < 3]
## [1] 1 2
# Easy way of work with parts of the data
data <- data.frame(x = c(0, 1, 3, 3, 0), y = 1:5)
data
## x y
## 1 0 1
## 2 1 2
## 3 3 3
## 4 3 4
## 5 0 5
# Data such that x is zero
data0 <- data[data$x == 0, ]
data0
## x y
## 1 0 1
## 5 0 5
# Data such that x is larger than 2
data2 <- data[data$x > 2, ]
data2
## x y
## 3 3 3
## 4 3 4
# In an example
iris$Sepal.Width[iris$Sepal.Width > 3]
## [1] 3.5 3.2 3.1 3.6 3.9 3.4 3.4 3.1 3.7 3.4 4.0 4.4 3.9 3.5 3.8 3.8 3.4 3.7 3.6
## [20] 3.3 3.4 3.4 3.5 3.4 3.2 3.1 3.4 4.1 4.2 3.1 3.2 3.5 3.6 3.4 3.5 3.2 3.5 3.8
## [39] 3.8 3.2 3.7 3.3 3.2 3.2 3.1 3.3 3.1 3.2 3.4 3.1 3.3 3.6 3.2 3.2 3.8 3.2 3.3
## [58] 3.2 3.8 3.4 3.1 3.1 3.1 3.1 3.2 3.3 3.4
# Problem - what happened?
data[x > 2, ]
## x y
## 3 3 3
## 4 3 4
## 5 0 5
# In an example
summary(iris)
## Sepal.Length Sepal.Width Petal.Length Petal.Width
## Min. :4.300 Min. :2.000 Min. :1.000 Min. :0.100
## 1st Qu.:5.100 1st Qu.:2.800 1st Qu.:1.600 1st Qu.:0.300
## Median :5.800 Median :3.000 Median :4.350 Median :1.300
## Mean :5.843 Mean :3.057 Mean :3.758 Mean :1.199
## 3rd Qu.:6.400 3rd Qu.:3.300 3rd Qu.:5.100 3rd Qu.:1.800
## Max. :7.900 Max. :4.400 Max. :6.900 Max. :2.500
## Species
## setosa :50
## versicolor:50
## virginica :50
##
##
##
summary(iris[iris$Sepal.Width > 3, ])
## Sepal.Length Sepal.Width Petal.Length Petal.Width
## Min. :4.400 Min. :3.100 Min. :1.000 Min. :0.1000
## 1st Qu.:5.000 1st Qu.:3.200 1st Qu.:1.450 1st Qu.:0.2000
## Median :5.400 Median :3.400 Median :1.600 Median :0.4000
## Mean :5.684 Mean :3.434 Mean :2.934 Mean :0.9075
## 3rd Qu.:6.400 3rd Qu.:3.600 3rd Qu.:5.000 3rd Qu.:1.8000
## Max. :7.900 Max. :4.400 Max. :6.700 Max. :2.5000
## Species
## setosa :42
## versicolor: 8
## virginica :17
##
##
##
# On the factor variable only makes sense == and !=
summary(iris[iris$Species == "setosa", ])
## Sepal.Length Sepal.Width Petal.Length Petal.Width
## Min. :4.300 Min. :2.300 Min. :1.000 Min. :0.100
## 1st Qu.:4.800 1st Qu.:3.200 1st Qu.:1.400 1st Qu.:0.200
## Median :5.000 Median :3.400 Median :1.500 Median :0.200
## Mean :5.006 Mean :3.428 Mean :1.462 Mean :0.246
## 3rd Qu.:5.200 3rd Qu.:3.675 3rd Qu.:1.575 3rd Qu.:0.300
## Max. :5.800 Max. :4.400 Max. :1.900 Max. :0.600
## Species
## setosa :50
## versicolor: 0
## virginica : 0
##
##
##
# Subset argument in lm
lm(Sepal.Width ~ Petal.Length, data = iris, subset = Sepal.Width > 3)
##
## Call:
## lm(formula = Sepal.Width ~ Petal.Length, data = iris, subset = Sepal.Width >
## 3)
##
## Coefficients:
## (Intercept) Petal.Length
## 3.59439 -0.05455
lm(Sepal.Width ~ Petal.Length, data = iris, subset = iris$Sepal.Width > 3)
##
## Call:
## lm(formula = Sepal.Width ~ Petal.Length, data = iris, subset = iris$Sepal.Width >
## 3)
##
## Coefficients:
## (Intercept) Petal.Length
## 3.59439 -0.05455
# Both iris$Sepal.Width and Sepal.Width in subset are fine: data = iris
# tells R to look for Sepal.Width in the iris dataset
# Same thing for the subset field in R Commander's menus
# AND operator &
TRUE & TRUE
## [1] TRUE
TRUE & FALSE
## [1] FALSE
FALSE & FALSE
## [1] FALSE
# OR operator |
TRUE | TRUE
## [1] TRUE
TRUE | FALSE
## [1] TRUE
FALSE | FALSE
## [1] FALSE
# Both operators are useful for checking for ranges of data
y
## [1] 0 3 1 5 2
index1 <- (y <= 3) & (y > 0)
y[index1]
## [1] 3 1 2
index2 <- (y < 2) | (y > 4)
y[index2]
## [1] 0 1 5
# In an example
summary(iris[iris$Sepal.Width > 3 & iris$Sepal.Width < 3.5, ])
## Sepal.Length Sepal.Width Petal.Length Petal.Width
## Min. :4.400 Min. :3.100 Min. :1.200 Min. :0.100
## 1st Qu.:4.925 1st Qu.:3.125 1st Qu.:1.500 1st Qu.:0.200
## Median :5.950 Median :3.200 Median :4.450 Median :1.400
## Mean :5.781 Mean :3.245 Mean :3.460 Mean :1.145
## 3rd Qu.:6.700 3rd Qu.:3.400 3rd Qu.:5.375 3rd Qu.:2.075
## Max. :7.200 Max. :3.400 Max. :6.000 Max. :2.500
## Species
## setosa :20
## versicolor: 8
## virginica :14
##
##
##
Do the following for the iris dataset:
-
Compute the subset corresponding to
Petal.Lengtheither smaller than1.5or larger than2. Save this dataset asirisPetal. -
Compute and summarize a linear regression of
Sepal.WidthintoPetal.Width + Petal.Lengthfor the datasetirisPetal. What is the \(R^2\)? (Solution:0.101) -
Check that the previous model is the same as regressing
Sepal.WidthintoPetal.Width + Petal.Lengthfor the datasetiriswith the appropriatesubsetexpression. -
Compute the variance for
Petal.WidthwhenPetal.Widthis smaller or equal that1.5and larger than0.3. (Solution:0.1266541)
4.1.4 Plotting functions
# plot is the main function for plotting in R
# It has a different behavior depending on the kind of object that it receives
# For example, for a regression model, it produces diagnostic plots
mod <- lm(Sepal.Width ~ Sepal.Length, data = iris)
plot(mod, 1)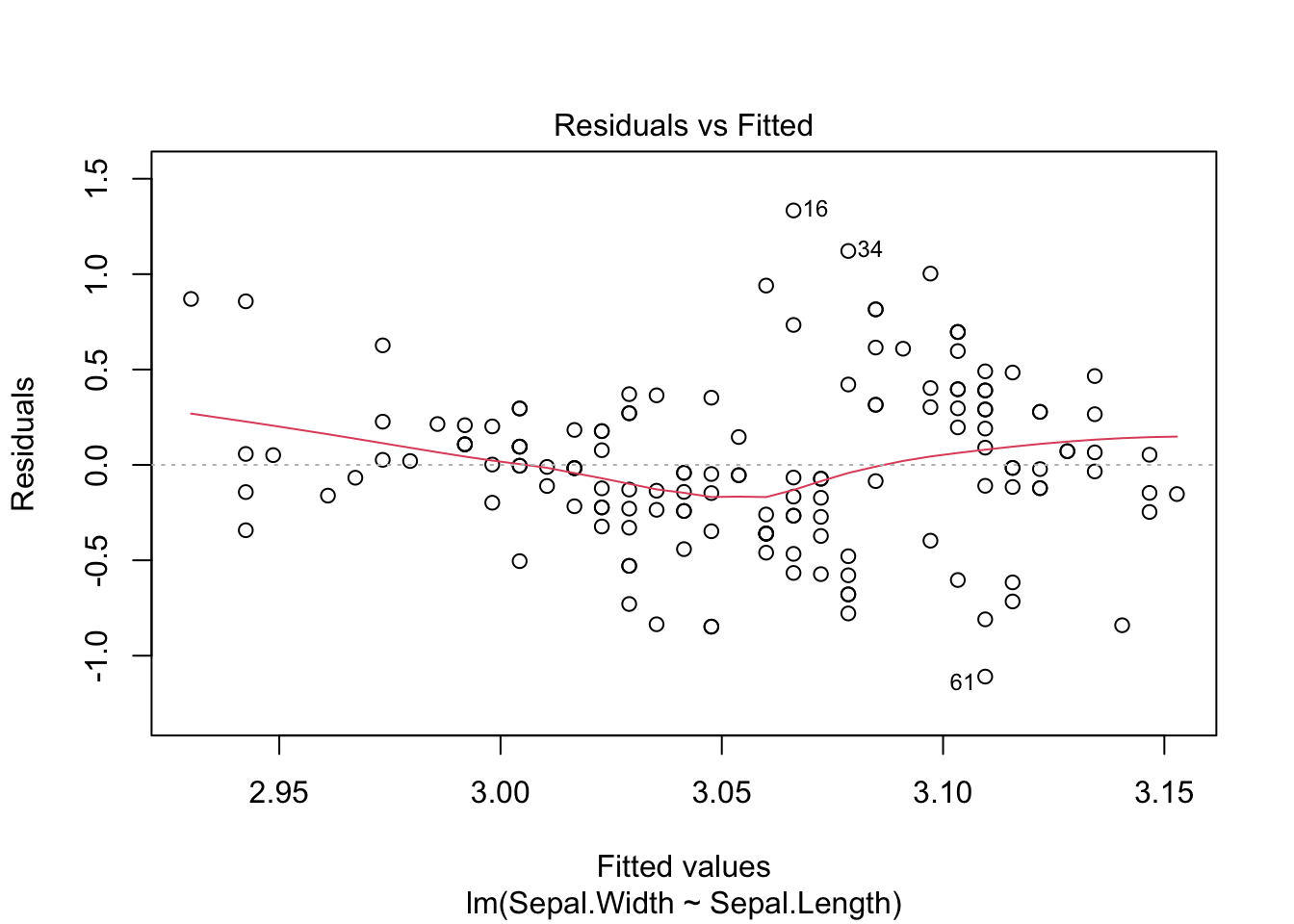
# How to plot some data
plot(iris$Sepal.Length, iris$Sepal.Width, main = "Sepal.Length vs Sepal.Width")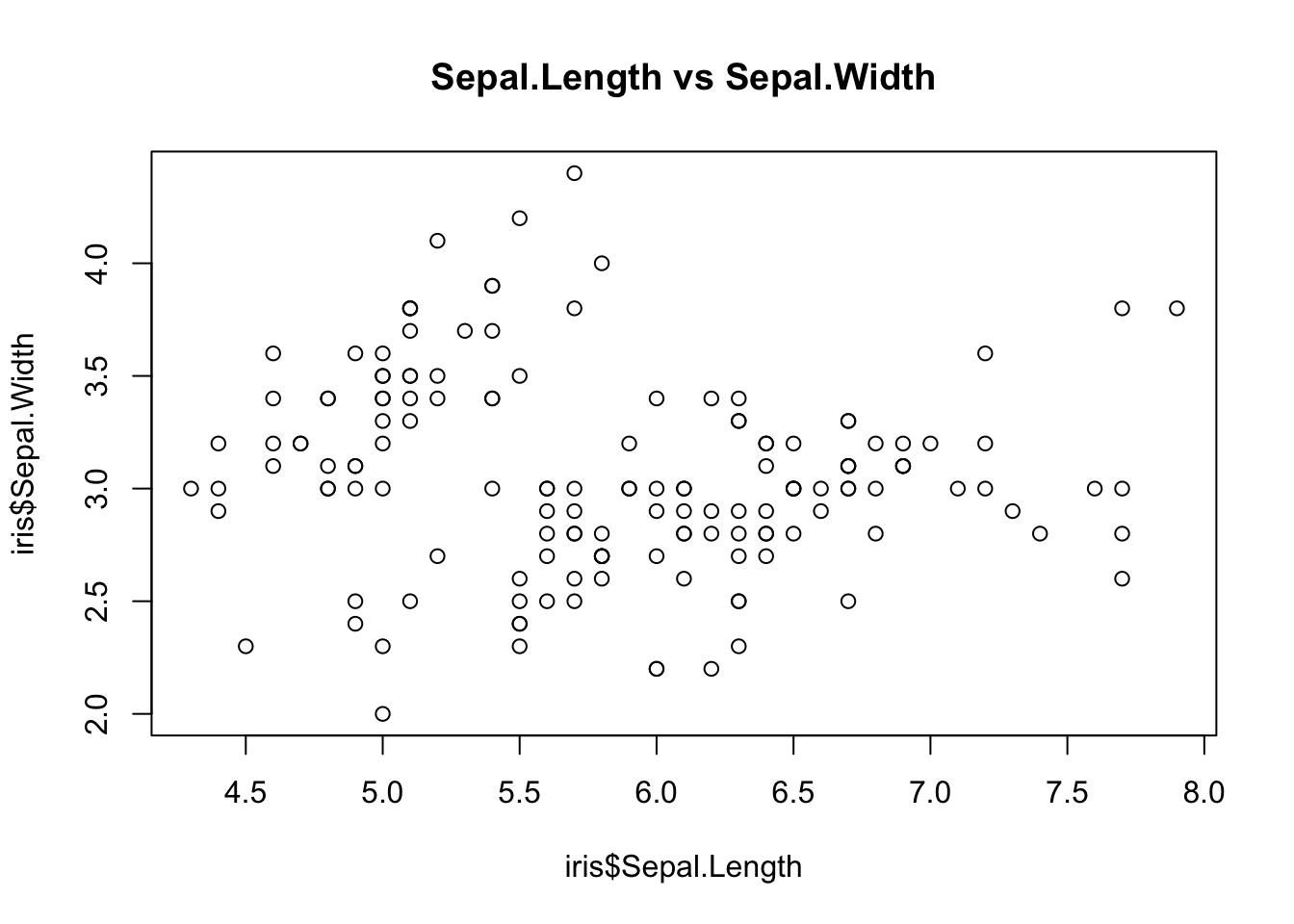
# Change the axis limits
plot(iris$Sepal.Length, iris$Sepal.Width, xlim = c(0, 10), ylim = c(0, 10))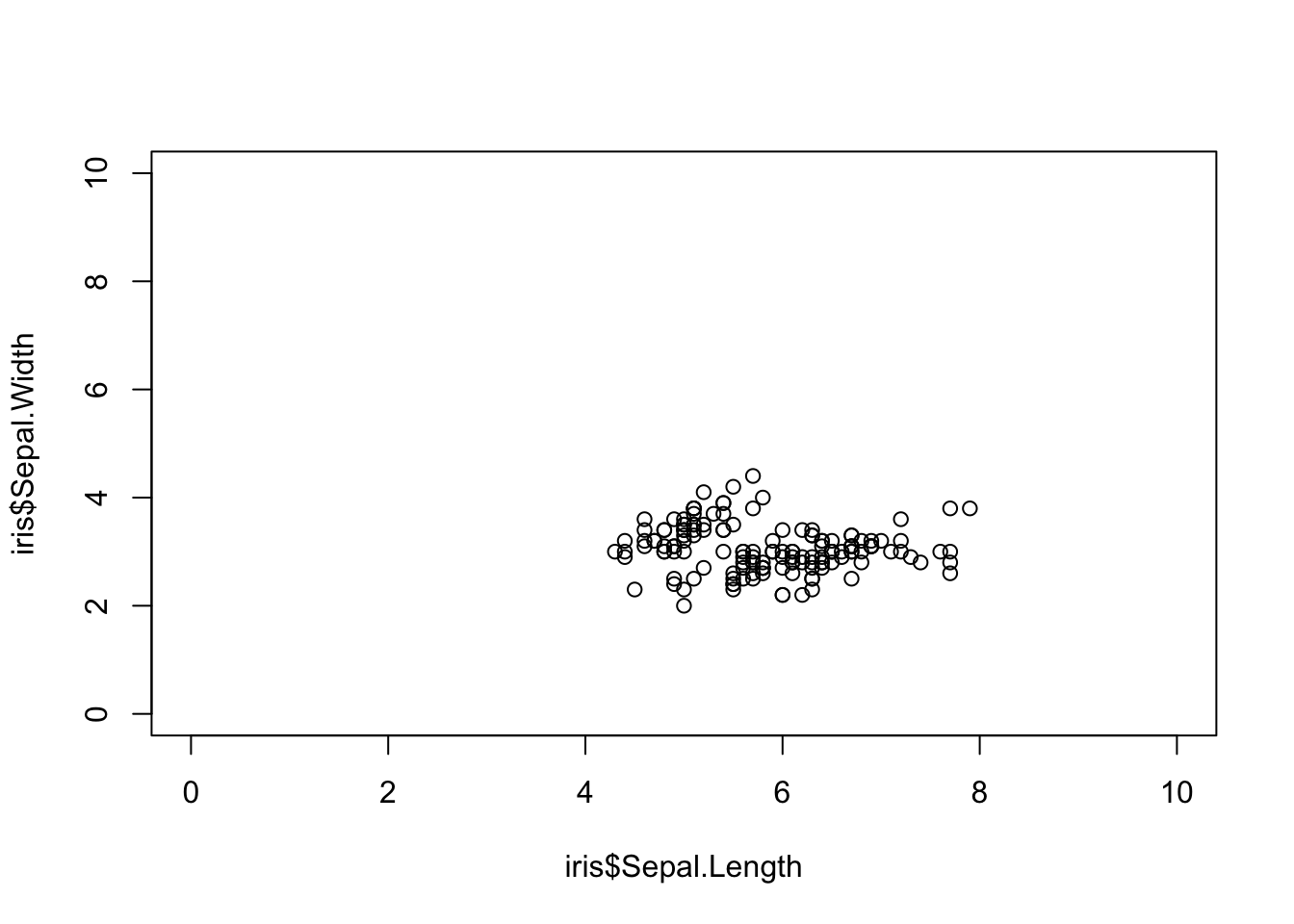
# How to plot a curve (a parabola)
x <- seq(-1, 1, l = 50)
y <- x^2
plot(x, y)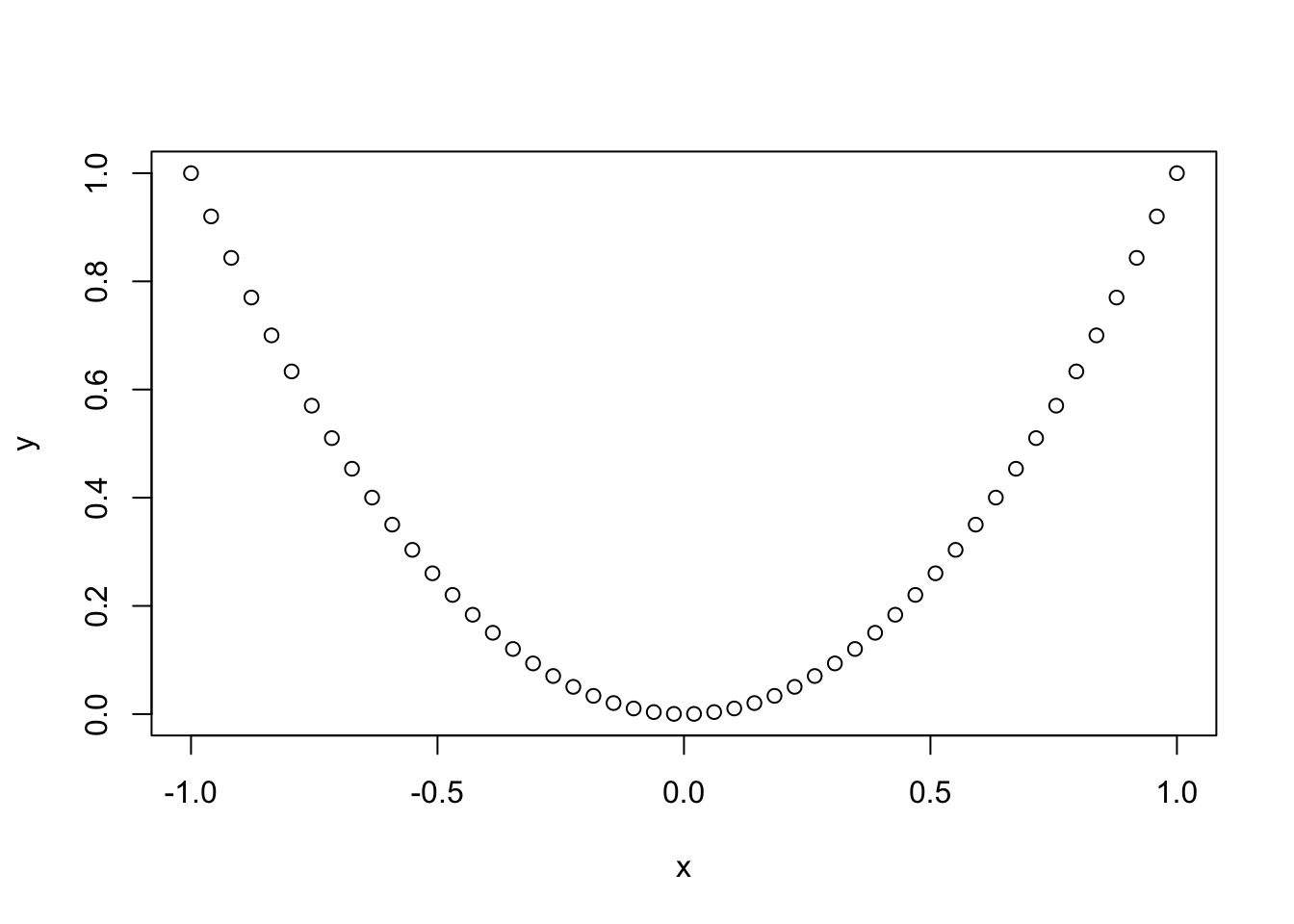
plot(x, y, main = "A dotted parabola")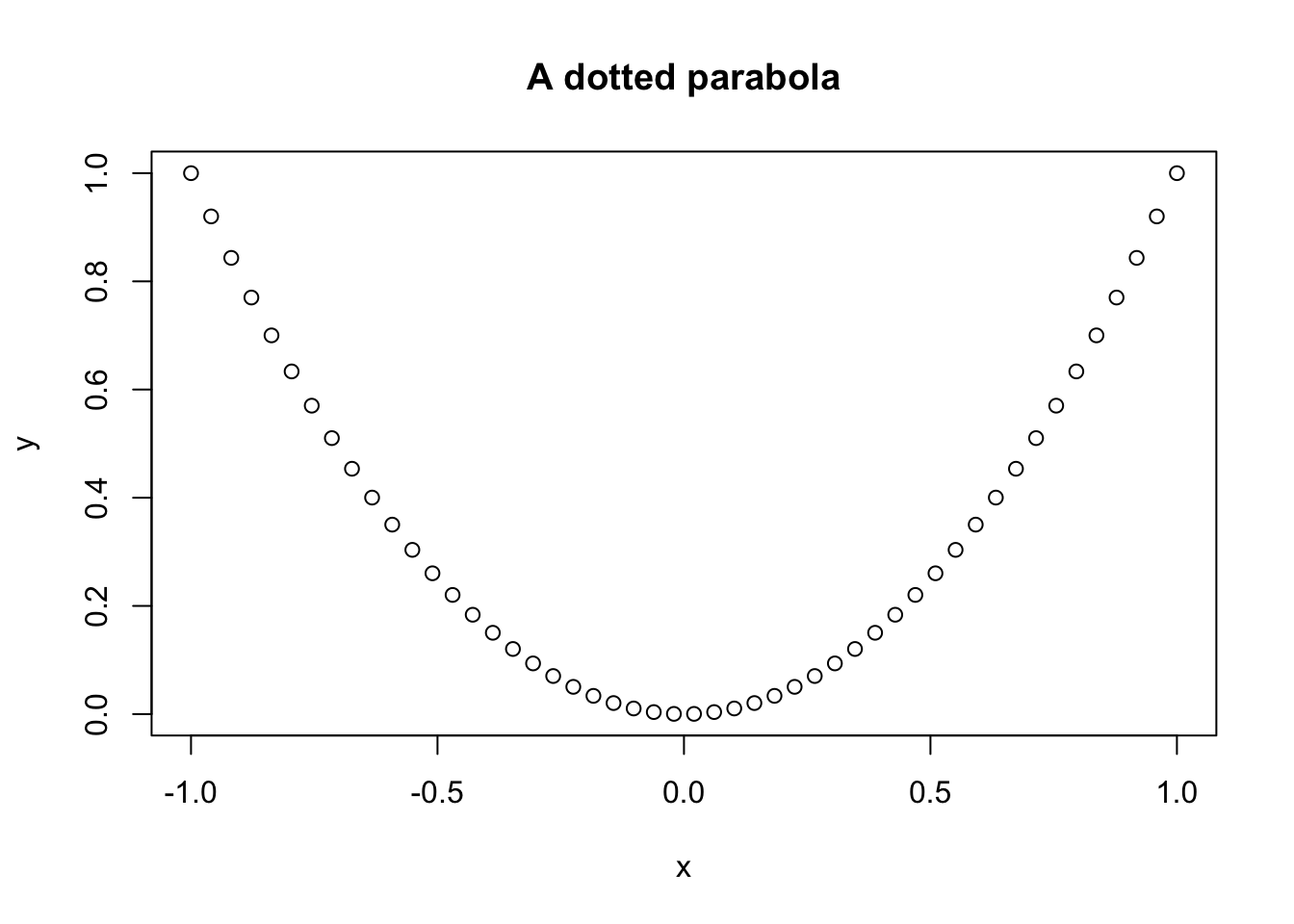
plot(x, y, main = "A parabola", type = "l")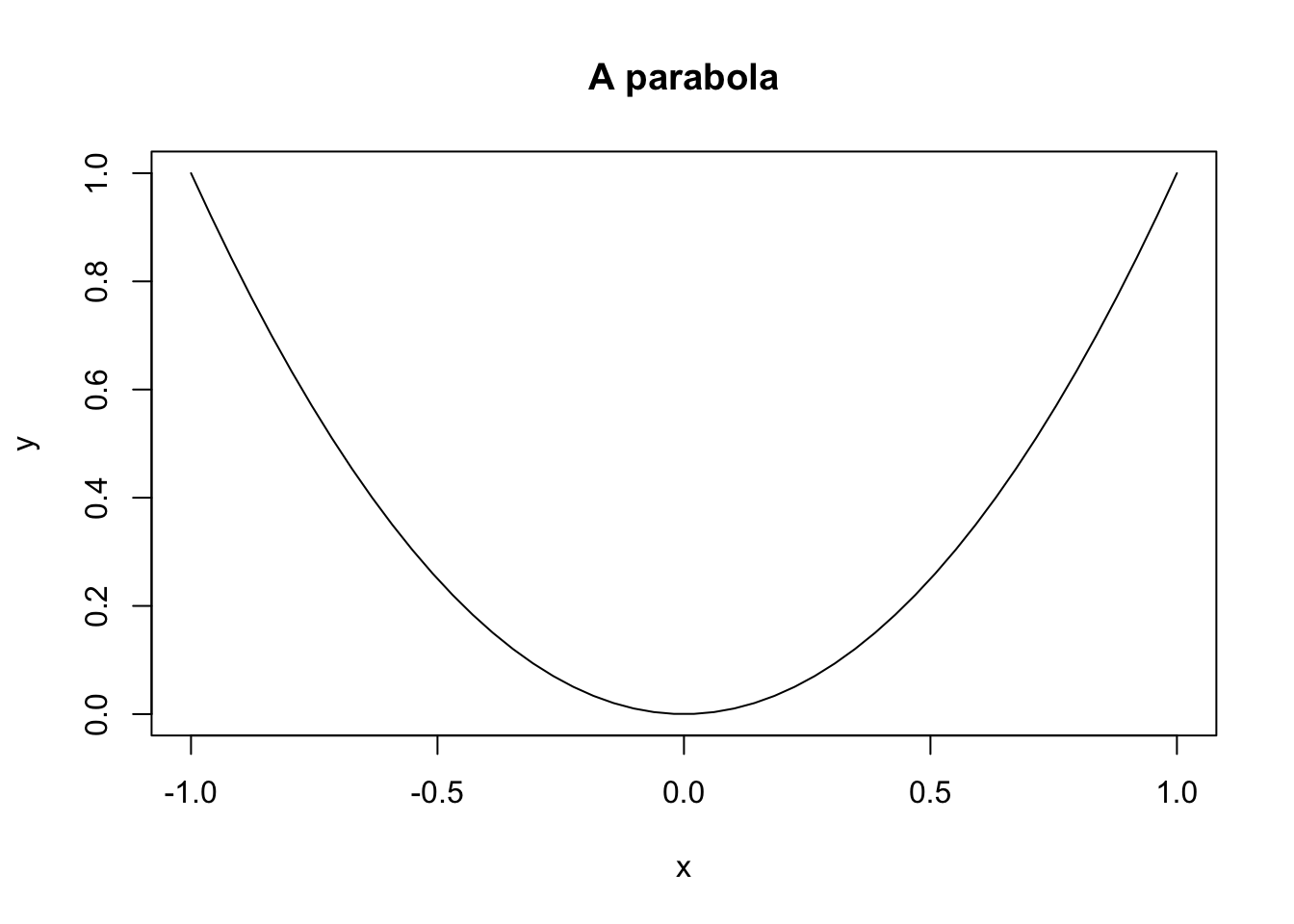
plot(x, y, main = "A red and thick parabola", type = "l", col = "red", lwd = 3)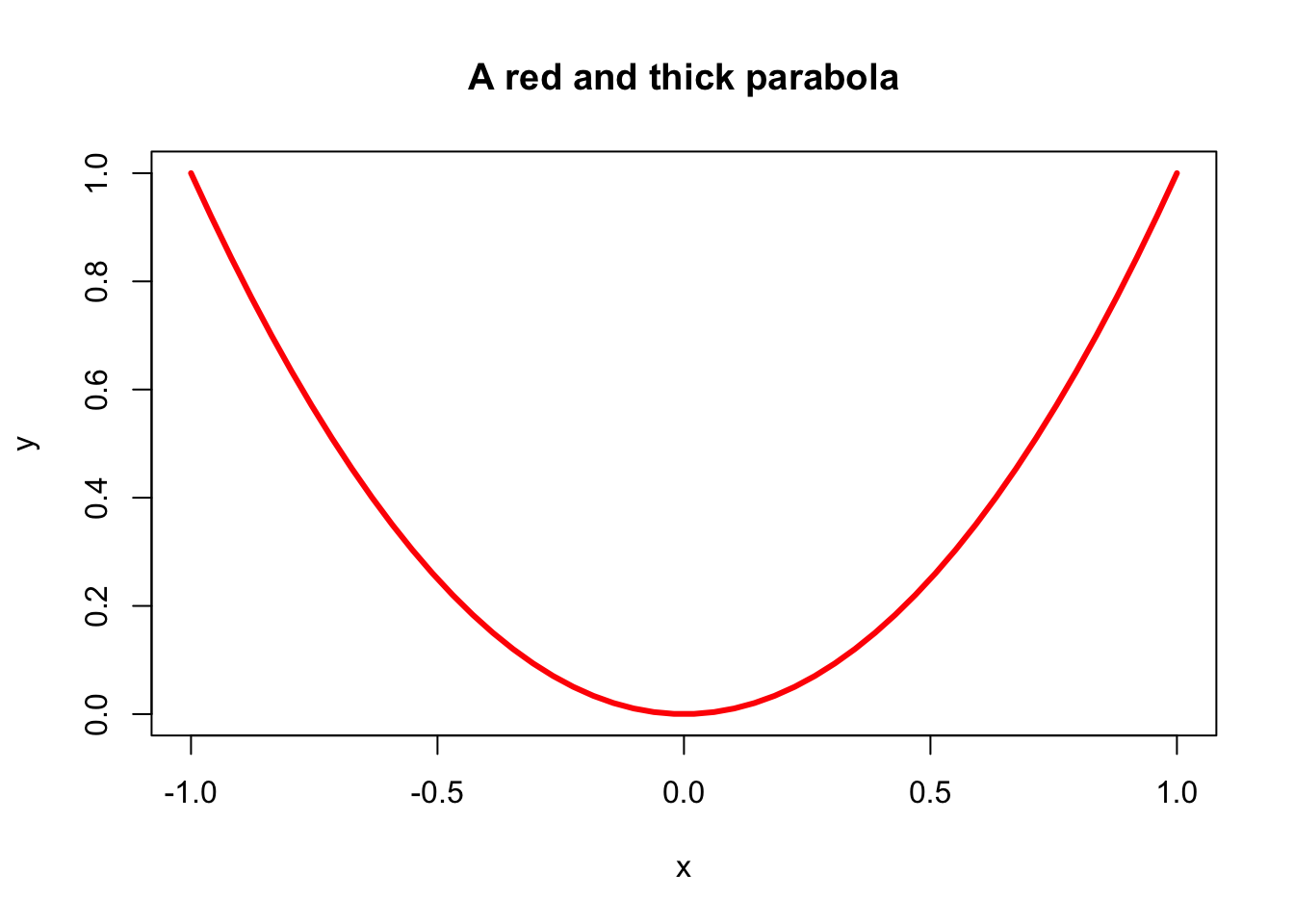
# Plotting a more complicated curve between -pi and pi
x <- seq(-pi, pi, l = 50)
y <- (2 + sin(10 * x)) * x^2
plot(x, y, type = "l") # Kind of rough...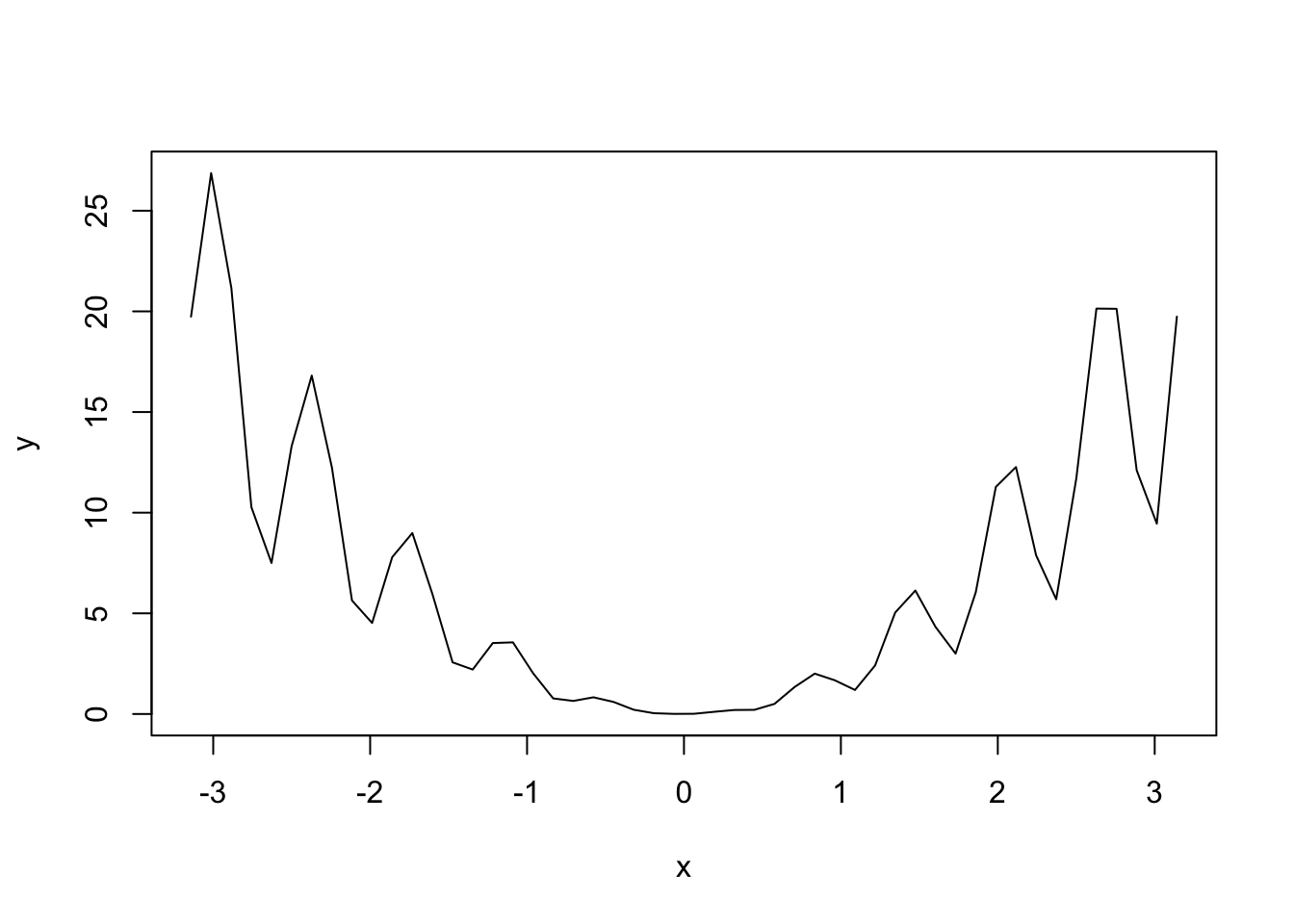
# More detailed plot
x <- seq(-pi, pi, l = 500)
y <- (2 + sin(10 * x)) * x^2
plot(x, y, type = "l")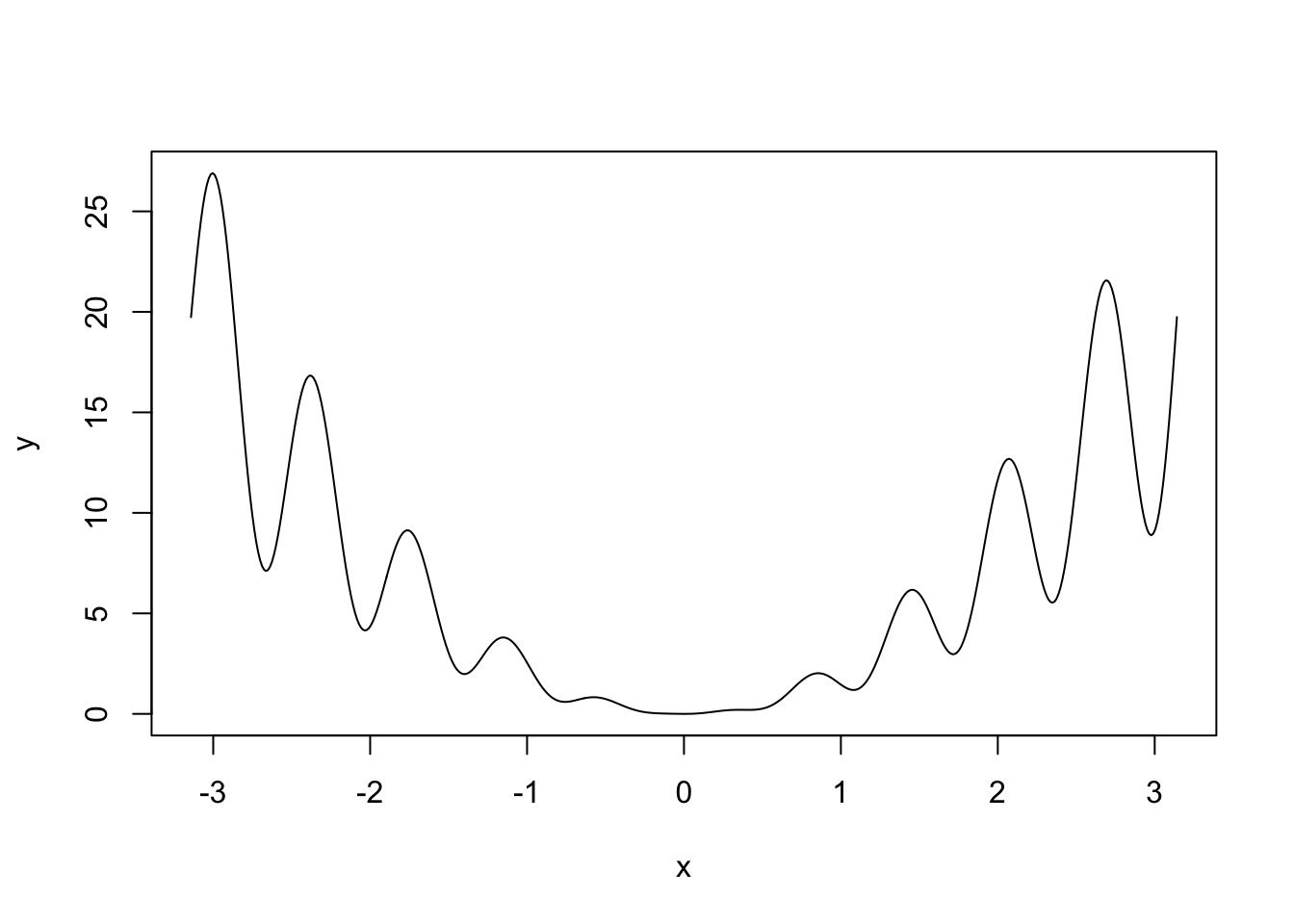
# Remember that we are joining points for creating a curve!
# For more options in the plot customization see
?plot
## Help on topic 'plot' was found in the following packages:
##
## Package Library
## base /Library/Frameworks/R.framework/Resources/library
## graphics /Library/Frameworks/R.framework/Versions/4.1-arm64/Resources/library
##
##
## Using the first match ...
?par
# plot is a first level plotting function. That means that whenever is called,
# it creates a new plot. If we want to add information to an existing plot, we
# have to use a second level plotting function such as points, lines or abline
plot(x, y) # Create a plot
lines(x, x^2, col = "red") # Add lines
points(x, y + 10, col = "blue") # Add points
abline(a = 5, b = 1, col = "orange", lwd = 2) # Add a straight line y = a + b * x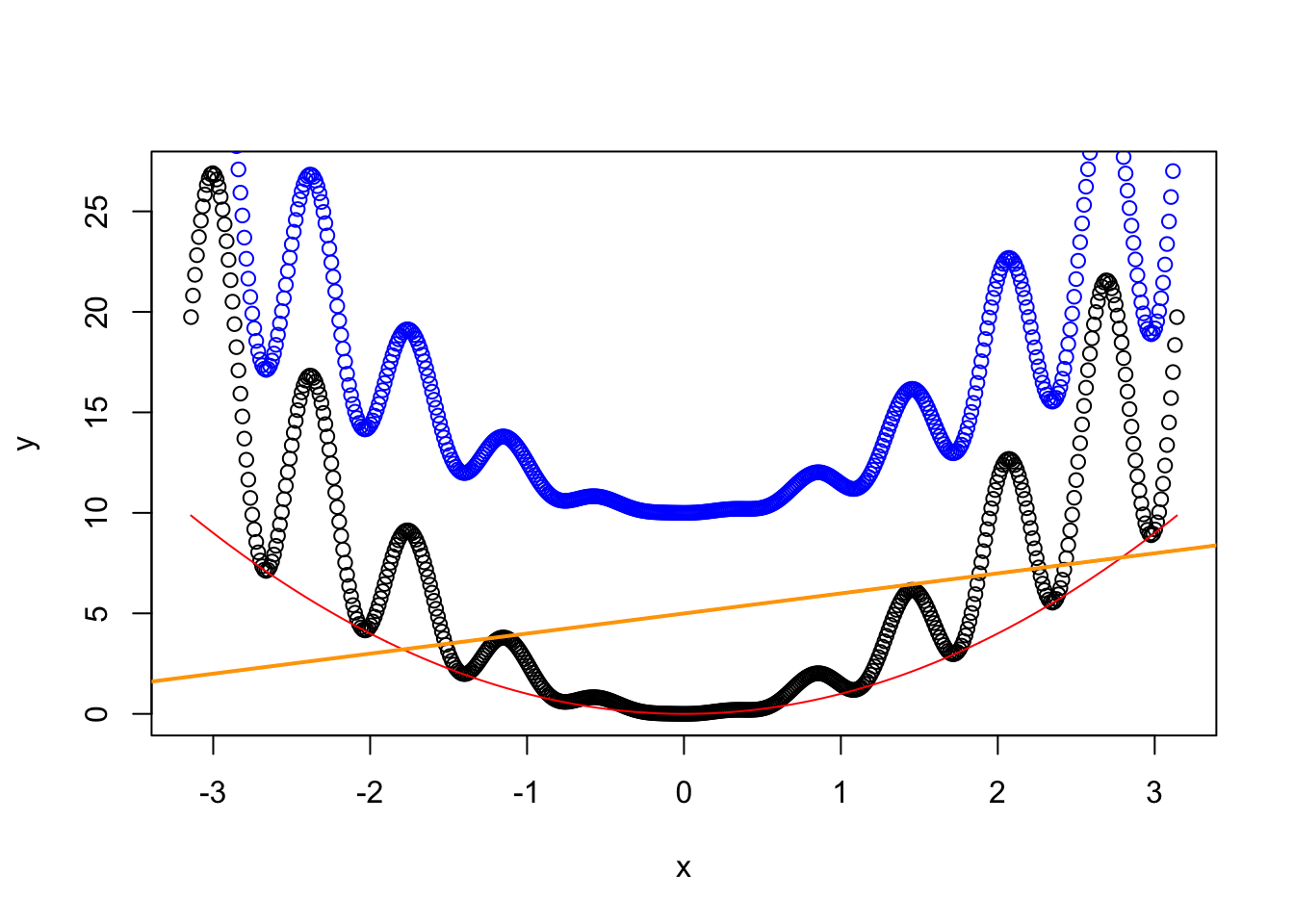
4.1.5 Distributions
The operations on distributions described here are implemented in R Commander through the menu 'Distributions', but is convenient for you to grasp how are they working.
# R allows to sample [r], compute density/probability mass [d],
# compute distribution function [p] and compute quantiles [q] for several
# continuous and discrete distributions. The format employed is [rdpq]name,
# where name stands for:
# - norm -> Normal
# - unif -> Uniform
# - exp -> Exponential
# - t -> Student's t
# - f -> Snedecor's F
# - chisq -> Chi squared
# - pois -> Poisson
# - binom -> Binomial
# More distributions:
?Distributions
# Sampling from a Normal - 100 random points from a N(0, 1)
rnorm(n = 10, mean = 0, sd = 1)
## [1] -1.8367426 -1.3366952 -0.4906582 1.0215158 0.1637865 2.5039127
## [7] 1.3113124 -0.1352548 0.1846896 -0.6373963
# If you want to have always the same result, set the seed of the random number
# generator
set.seed(45678)
rnorm(n = 10, mean = 0, sd = 1)
## [1] 1.4404800 -0.7195761 0.6709784 -0.4219485 0.3782196 -1.6665864
## [7] -0.5082030 0.4433822 -1.7993868 -0.6179521
# Plotting the density of a N(0, 1) - the Gauss bell
x <- seq(-4, 4, l = 100)
y <- dnorm(x = x, mean = 0, sd = 1)
plot(x, y, type = "l")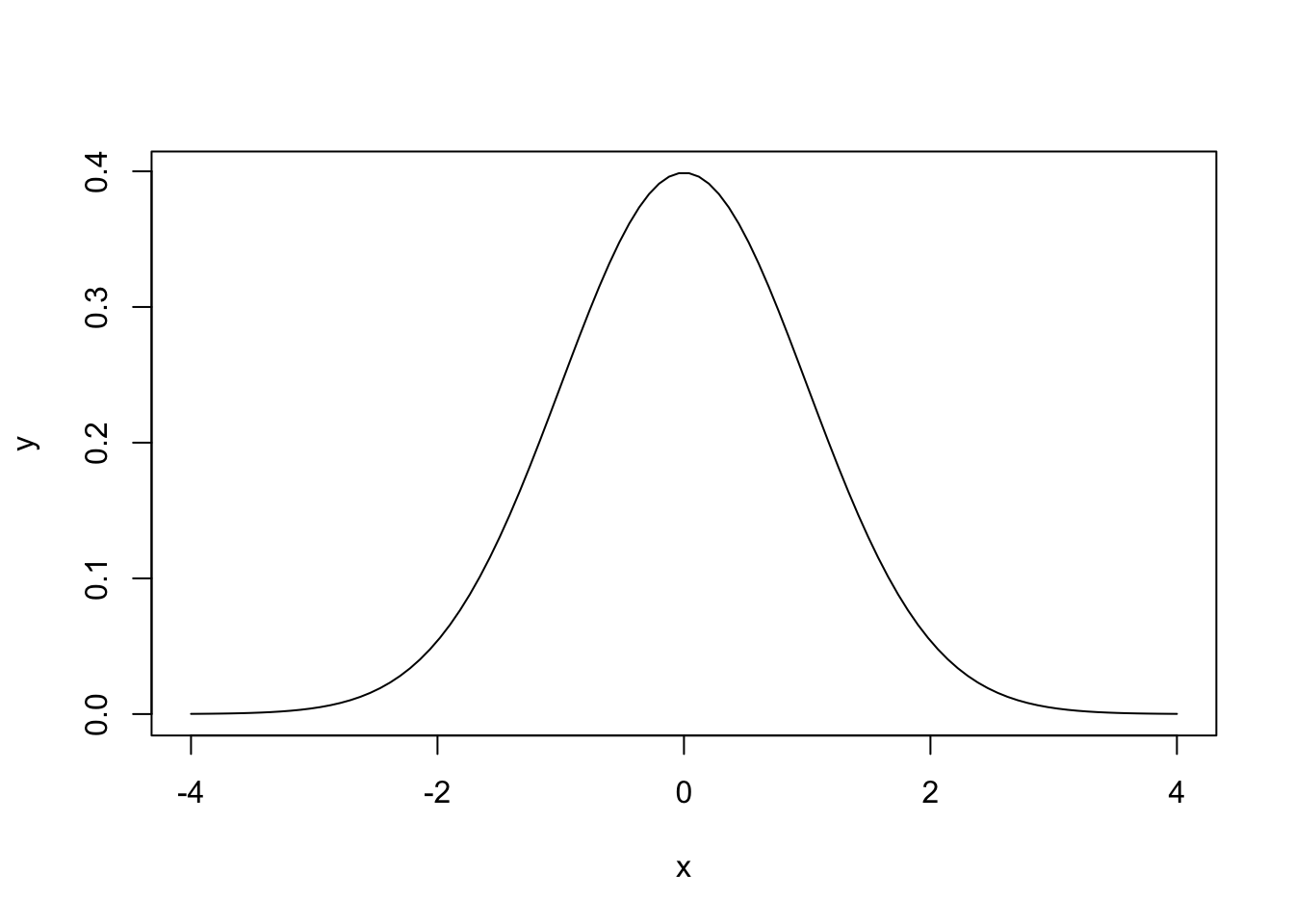
# Plotting the distribution function of a N(0, 1)
x <- seq(-4, 4, l = 100)
y <- pnorm(q = x, mean = 0, sd = 1)
plot(x, y, type = "l")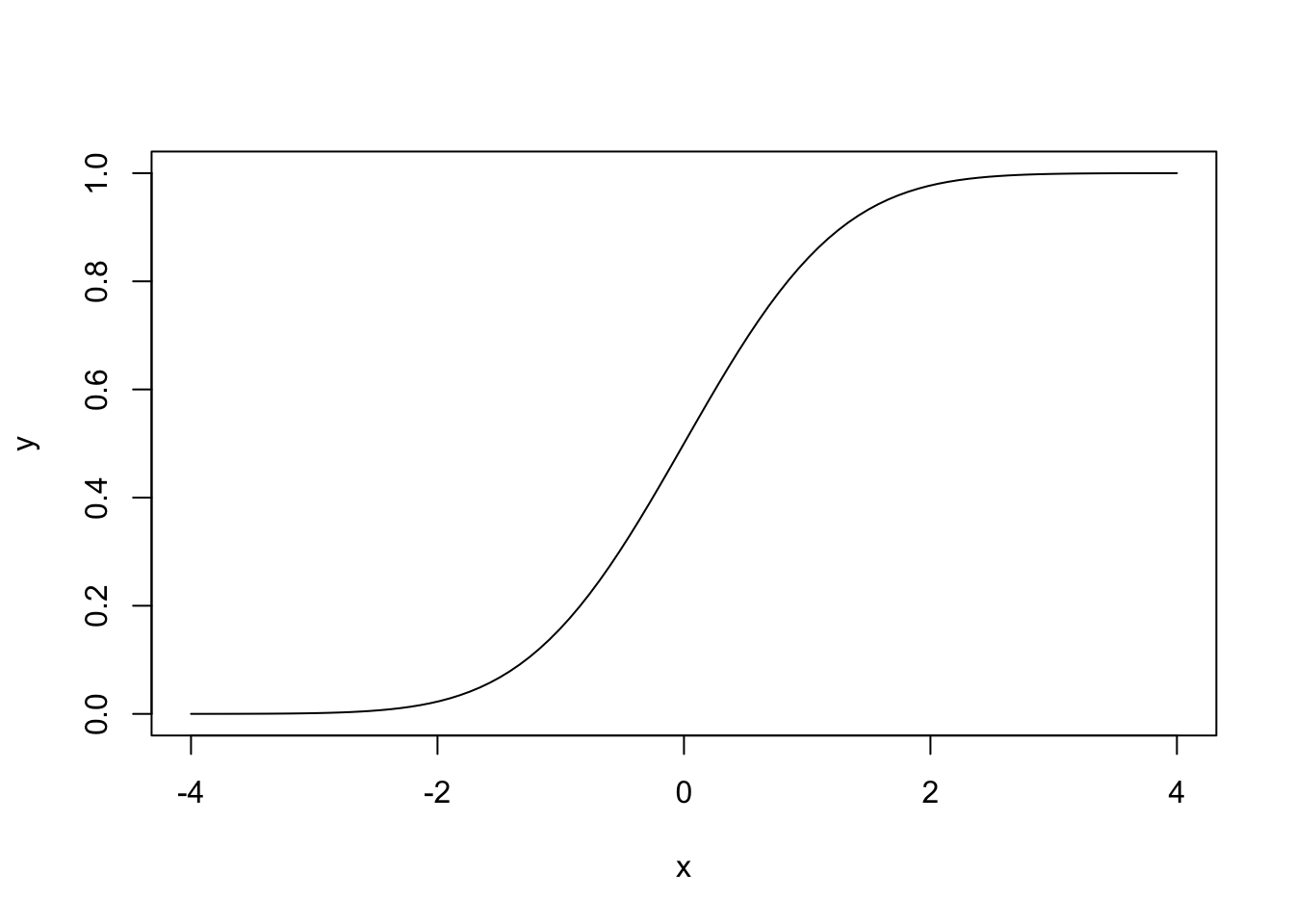
# Computing the 95% quantile for a N(0, 1)
qnorm(p = 0.95, mean = 0, sd = 1)
## [1] 1.644854
# All distributions have the same syntax: rname(n,...), dname(x,...), dname(p,...)
# and qname(p,...), but the parameters in ... change. Look them in ?Distributions
# For example, here is the same for the uniform distribution
# Sampling from a U(0, 1)
set.seed(45678)
runif(n = 10, min = 0, max = 1)
## [1] 0.9251342 0.3339988 0.2358930 0.3366312 0.7488829 0.9327177 0.3365313
## [8] 0.2245505 0.6473663 0.0807549
# Plotting the density of a U(0, 1)
x <- seq(-2, 2, l = 100)
y <- dunif(x = x, min = 0, max = 1)
plot(x, y, type = "l")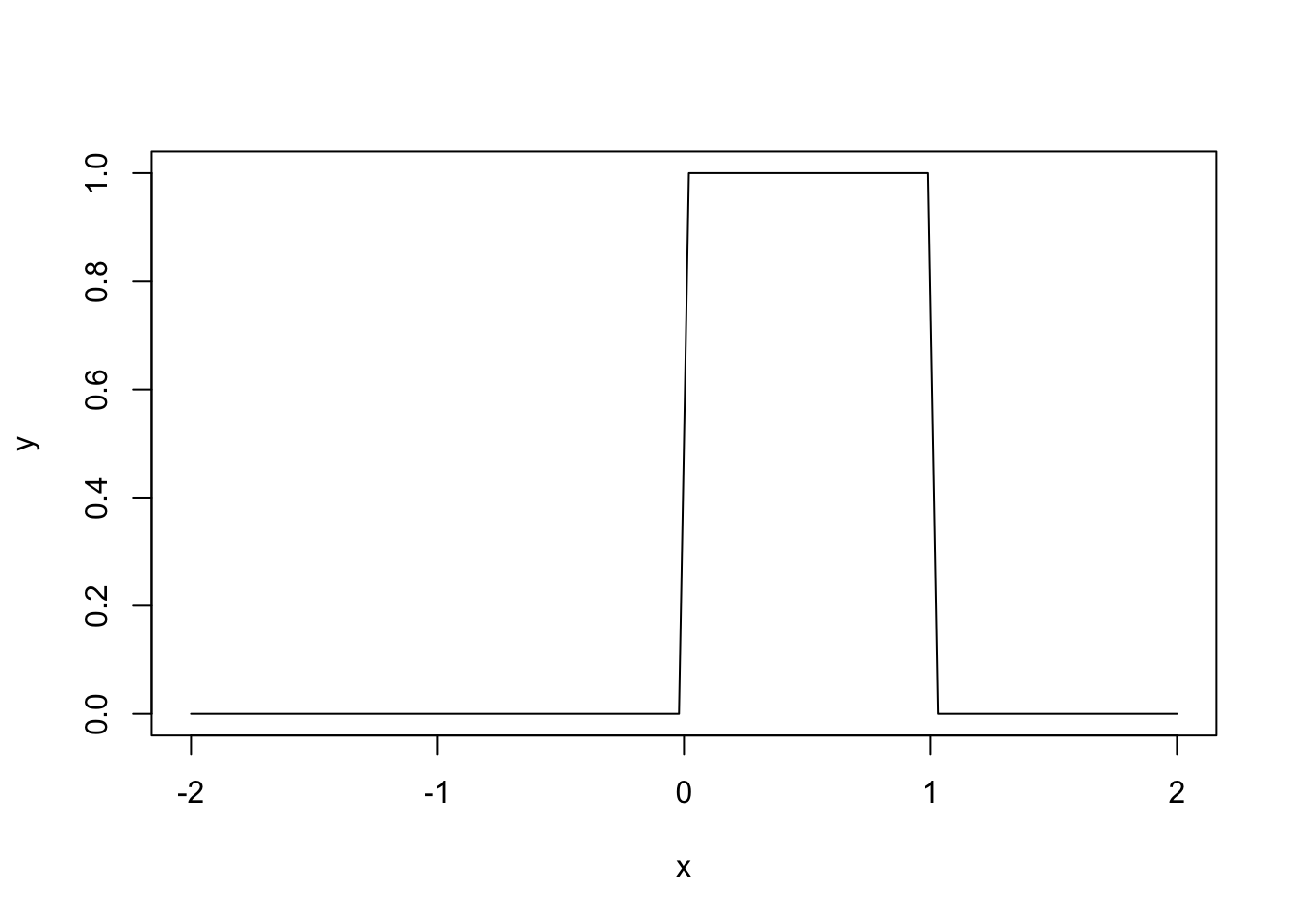
# Computing the 95% quantile for a U(0, 1)
qunif(p = 0.95, min = 0, max = 1)
## [1] 0.95
# Sampling from a Bi(10, 0.5)
set.seed(45678)
samp <- rbinom(n = 200, size = 10, prob = 0.5)
table(samp) / 200
## samp
## 1 2 3 4 5 6 7 8 9
## 0.010 0.060 0.115 0.220 0.210 0.215 0.115 0.045 0.010
# Plotting the probability mass of a Bi(10, 0.5)
x <- 0:10
y <- dbinom(x = x, size = 10, prob = 0.5)
plot(x, y, type = "h") # Vertical bars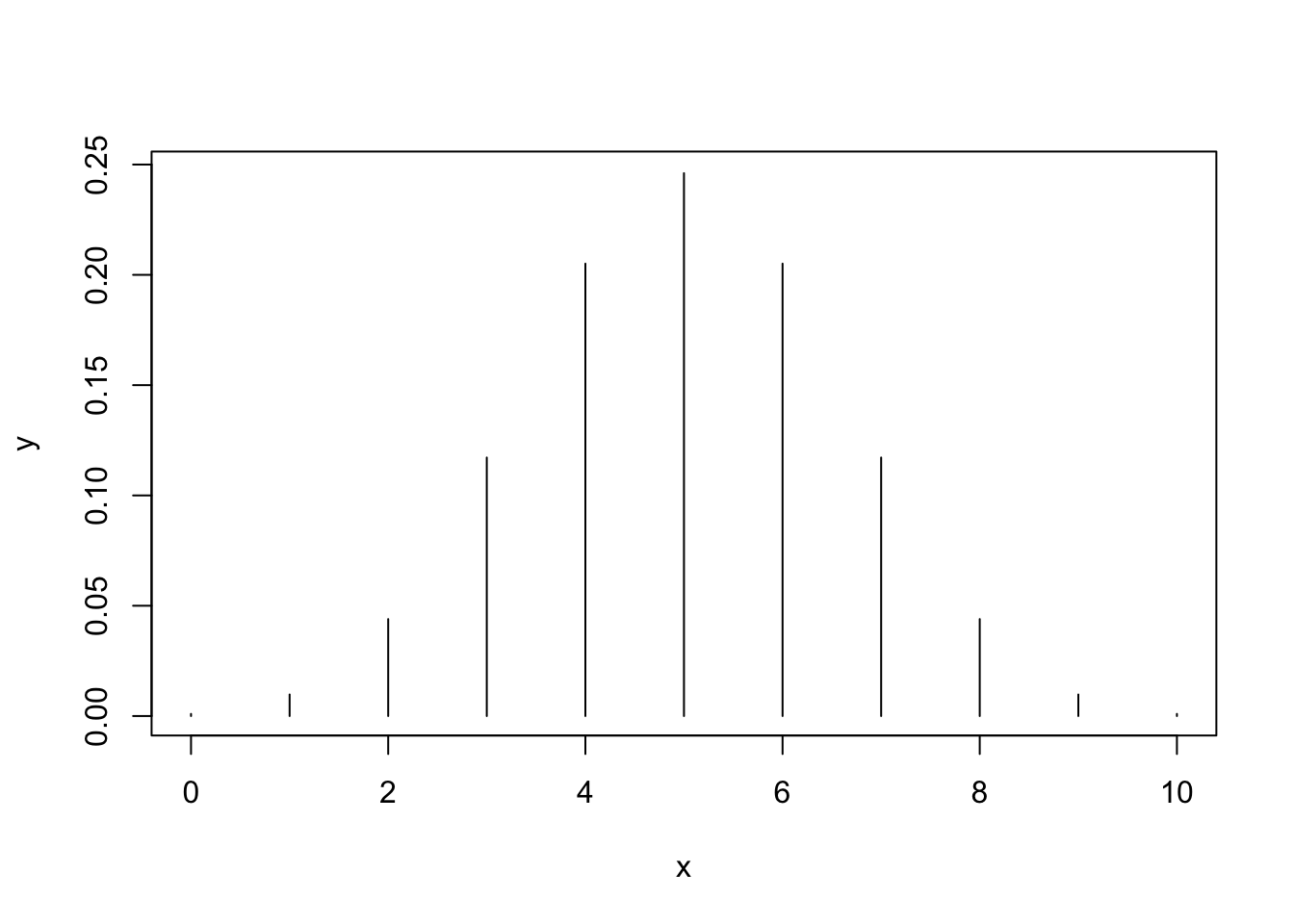
# Plotting the distribution function of a Bi(10, 0.5)
x <- 0:10
y <- pbinom(q = x, size = 10, prob = 0.5)
plot(x, y, type = "h")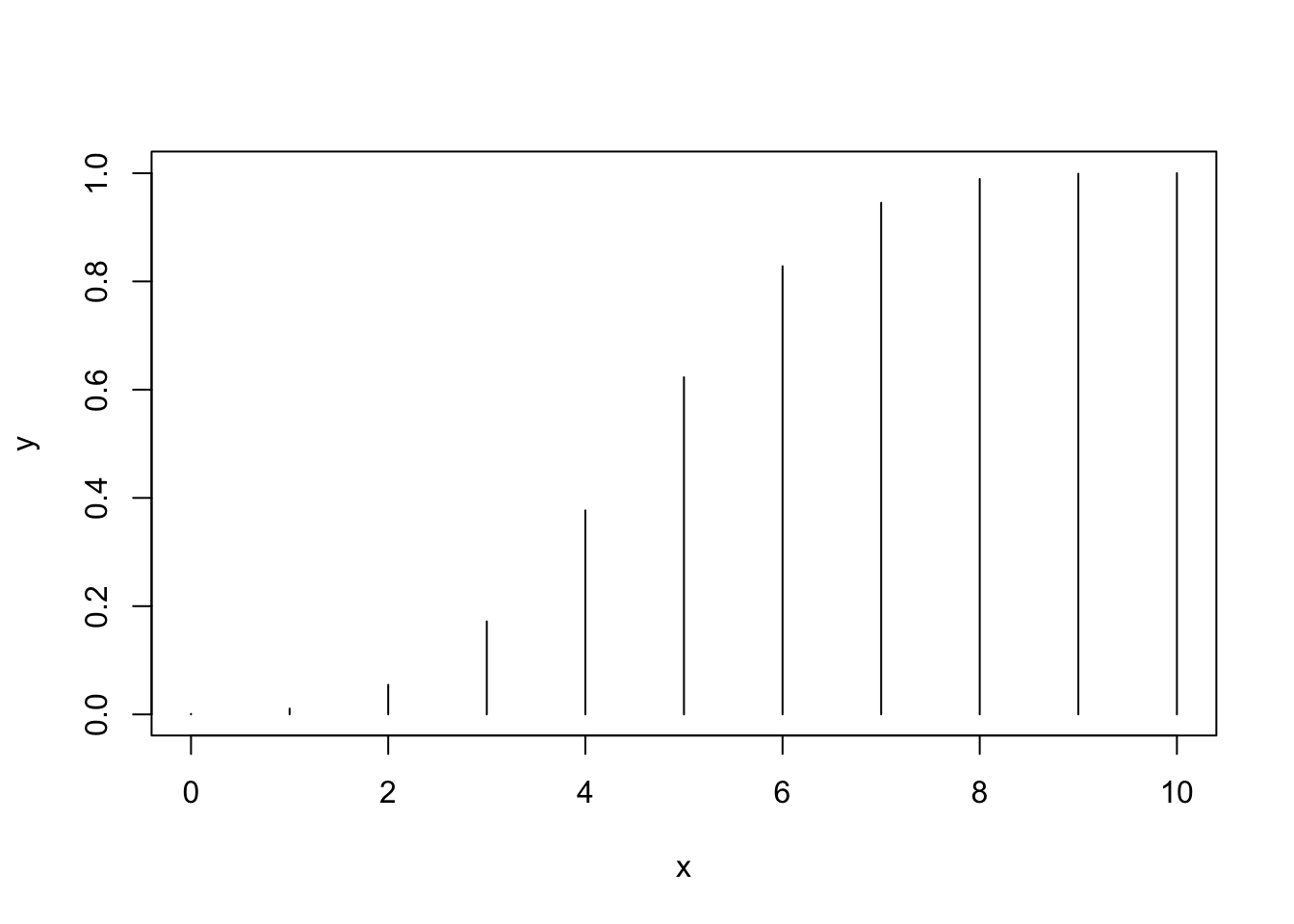
Do the following:
-
Compute the 90%, 95% and 99% quantiles of a \(F\) distribution with
df1 = 1anddf2 = 5. (Answer:c(4.060420, 6.607891, 16.258177)) - Plot the distribution function of a \(U(0,1)\). Does it make sense with its density function?
-
Sample 100 points from a Poisson with
lambda = 5. - Sample 100 points from a \(U(-1,1)\) and compute its mean.
-
Plot the density of a \(t\) distribution with
df = 1(use a sequence spanning from-4to4). Add lines of different colors with the densities fordf = 5,df = 10,df = 50anddf = 100. Do you see any pattern?
4.1.6 Defining functions
# A function is a way of encapsulating a block of code so it can be reused easily
# They are useful for simplifying repetitive tasks and organize the analysis
# For example, in Section 3.7 we had to make use of simpleAnova for computing
# the simple ANOVA table in multiple regression.
# This is a silly function that takes x and y and returns its sum
add <- function(x, y) {
x + y
}
# Calling add - you need to run the definition of the function first!
add(1, 1)
## [1] 2
add(x = 1, y = 2)
## [1] 3
# A more complex function: computes a linear model and its posterior summary.
# Saves us a few keystrokes when computing a lm and a summary
lmSummary <- function(formula, data) {
model <- lm(formula = formula, data = data)
summary(model)
}
# Usage
lmSummary(Sepal.Length ~ Petal.Width, iris)
##
## Call:
## lm(formula = formula, data = data)
##
## Residuals:
## Min 1Q Median 3Q Max
## -1.38822 -0.29358 -0.04393 0.26429 1.34521
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 4.77763 0.07293 65.51 <2e-16 ***
## Petal.Width 0.88858 0.05137 17.30 <2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.478 on 148 degrees of freedom
## Multiple R-squared: 0.669, Adjusted R-squared: 0.6668
## F-statistic: 299.2 on 1 and 148 DF, p-value: < 2.2e-16
# Recall: there is no variable called model in the workspace.
# The function works on its own workspace!
model
##
## Call:
## lm(formula = medv ~ crim + lstat + zn + nox, data = Boston)
##
## Coefficients:
## (Intercept) crim lstat zn nox
## 30.93462 -0.08297 -0.90940 0.03493 5.42234
# Add a line to a plot
addLine <- function(x, beta0, beta1) {
lines(x, beta0 + beta1 * x, lwd = 2, col = 2)
}
# Usage
plot(x, y)
addLine(x, beta0 = 0.1, beta1 = 0)