4.9 Vectors
A vector is a collection of values. “Vector” means different things in different fields (mathematics, geometry, biology), but in R it is a fancy name for a collection of values. We call the individual values elements of the vector. It is one of the most common data structures you will work with in R.
We can make vectors with the function c( ), for example c(1,2,3). c
means “combine.” R is obsessed with vectors, in R even single numbers
are vectors of length one. Many things that can be done with a single
number can also be done with a vector. For example arithmetic can be
done on vectors as it can be on single numbers.
Let’s say that we have a group of patients in our clinic. We can store their names in a vector.
patients <- c("Maria", "John", "Ali", "Luis", "Mei" )
patients## [1] "Maria" "John" "Ali" "Luis" "Mei"If we later wanted to add a name, it’s easy to do so
patients <- c(patients, "Emma")
patients## [1] "Maria" "John" "Ali" "Luis" "Mei" "Emma"Maybe we also want to store the weights of these patients. Since these are their weights in pounds, we will call our object weight_lb
weight_lb <- c(122, 320, 217, 142, 174, 252)
weight_lb## [1] 122 320 217 142 174 252So far, we have created vectors of two different data types: character and numeric.
You can do arithmatic with numeric vectors. For example, let’s convert the weight of our patients in lbs to the the weight in kilograms by multiplying each weight in lbs by 2.2.
We could do this one by one:
122 / 2.2## [1] 55.45455320 / 2.2## [1] 145.4545217 / 2.2## [1] 98.63636etc.
But that would be a long and tedious process, especially if you had more than 6 patients.
Instead, let’s divide the vector by 2.2 and save that to a new object. We will call this object weight_kg
weight_kg <- weight_lb / 2.2
#you could also round the weight
weight_kg <- round((weight_lb / 2.2), digits = 2)
weight_kg## [1] 55.45 145.45 98.64 64.55 79.09 114.55We could use the mean() function to find out the mean weight of patients at our clinic.
mean(weight_lb)## [1] 204.5You can not do this with character vectors. Remember we used c() to add a value to our character vector.
patients + "Sue"## Error in patients + "Sue": non-numeric argument to binary operatorYou can combine two vectors together
Data Types
There are numerous data types. Some of the other most common data types you will encounter are numeric data, character data and logical data. Vectors of one data type only are called atomic vectors. Read more about vectors and data types in the book R for Data Science
Another common data type is logical data. Logical data is the valuesTRUE, FALSE, or NA
We want to record if our patients have been fully vaccinated. We will record this as TRUE if they have been, FALSE if they have not been, and NA if we do not have this information.
vax_status <- c(TRUE, TRUE, FALSE, NA, TRUE, FALSE)
vax_status## [1] TRUE TRUE FALSE NA TRUE FALSEAll vector types have a length property which you can determine with the length() function.
length(patients)## [1] 6Its helpful to think of the length of a vector as the number of elements in the vector.
You can always find out the data type of your vector with the class() function.
class(patients)## [1] "character"class(weight_lb)## [1] "numeric"class(vax_status)## [1] "logical"4.9.1 Missing Data
R also has many tools to help with missing data, a very common occurrence.
Suppose you tried to calculate the mean of a vector with some missing values (represented here with the logical NA. For example, what if we had failed to capture the weight of some of the patients at our clinic, so our weight vector looks as follows:
missing_wgt <- c(122, NA, 217, NA, 174, 252)
mean(missing_wgt)## [1] NAThe missing values cause an error, and the mean cannot be correctly calculated.
To get around this, you can use the argument na.rm = TRUE. This says to remove the NA values before attempting to perform the calculation.
mean(missing_wgt, na.rm = TRUE)## [1] 191.25For more on how to work with missing data, check out this Data Carpentry lesson
Factors
Another important type of vector is a factor. Factors are a way that R stores categorical variables. In a factor, the levels of a categorical value are mapped onto an vector of integers, much like if you were coding responses to a survey. So, it is important to be careful because factors look like character data, but need to be treated like numeric data. For more on factors check out the Software Carpentry lesson on R
4.9.2 Mixing types
We said above that vectors are supposed to have only one data type, but what happens if we mix multiple data types in one vector?
Sometimes the best way to understand R is to try some examples and see what it does.
Questions
Create a vector with some patient names and weights together. What happens? What if you combine names and vax status? All three?
Because vectors can only contain one type of thing, R chooses a lowest common denominator type of vector, a type that can contain everything we are trying to put in it. A different language might stop with an error, but R tries to soldier on as best it can. A number can be represented as a character string, but a character string can not be represented as a number, so when we try to put both in the same vector R converts everything to a character string.
4.9.3 Indexing and Subsetting vectors
Access elements of a vector with [ ], for example
patients[1]## [1] "Maria" patients[4]## [1] "Luis"You can also assign to a specific element of a vector.
patients[2] <- "Jon"
patients## [1] "Maria" "Jon" "Ali" "Luis" "Mei" "Emma"Can we use a vector to index another vector? Yes!
vaxInd <- c(1,2,5)
patients[vaxInd]## [1] "Maria" "Jon" "Mei" vax_patients <- patients[vaxInd]
vax_patients## [1] "Maria" "Jon" "Mei"We could equivalently have written:
patients[c(1, 3, 5)]## [1] "Maria" "Ali" "Mei"4.9.4 Data frames and tibbles
The other main data structure you are likely to work with is the data frame. Vectors are one dimensional data structures. A data frame is two dimensional (often called tabular or rectangular data). This is similar to data as you might be used to seeing it in a spreadsheet. You can think of a data frame as consisting of columns of vectors. As vectors,each column in a data frame is of one data type, but the data frame over all can hold multiple data types.
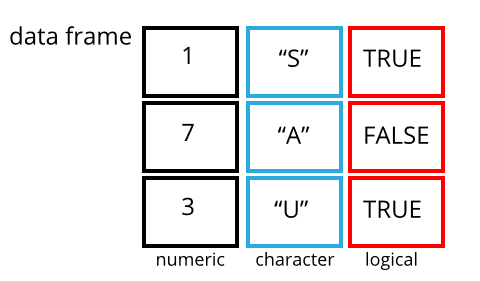
A tibble is a particular class of data frame which is common in the tidyverse family of packages. Tibbles are useful for their printing properties and because they are less likely try to change the data type of columns on import (e.g. from character to factor).
Since vectors form the columns of data frames, we can take the vectors we created for our patient data and combine them together in a data frame.
patient_data <- data.frame(patients, weight_lb, weight_kg, vax_status)
patient_data## patients weight_lb weight_kg vax_status
## 1 Maria 122 55.45 TRUE
## 2 Jon 320 145.45 TRUE
## 3 Ali 217 98.64 FALSE
## 4 Luis 142 64.55 NA
## 5 Mei 174 79.09 TRUE
## 6 Emma 252 114.55 FALSE