Chapter 19 Maximum Likelihood Estimation for the General Linear Model
Likelihood function: \(L(\theta\mid y) = f(y\mid \theta)\) is a real-valued function of \(\theta\) for a given value of \(y\).
Maximum likelihood estimatior: \(\hat\theta = \arg\max_{\theta} L(\theta\mid y)\)
Score function: \(\frac{\partial \ell(\theta\mid y)}{\partial \theta}\)
Score equation: \(\frac{\partial \ell(\theta\mid y)}{\partial \theta} = 0\)
- Example: \[ \begin{bmatrix} \hat\beta \\ n^{-1}(y-X\hat\beta)'(y-X\hat\beta)\end{bmatrix} \text{ is an MLE of }\theta = \begin{bmatrix} \beta \\ \theta \end{bmatrix} \] However, the MLE of \(\sigma^2\) is not an unbiased estimator, \(E(SSE/n) = \frac{n-r}{n}\sigma^2\)
Profiled Log Likelihood: example \(\ell^*(\gamma\mid y) = \ell(\begin{bmatrix} \hat\beta_{\Sigma} \\\ \gamma \end{bmatrix} \mid y)\) where we suppose \(y = X\beta + \epsilon\), \(\epsilon \sim N(0, \Sigma)\), \(\Sigma = \sigma^2\begin{bmatrix} 1 & \rho & \rho^2 \\ \rho & 1 & \rho \\\ \rho^2 & rho & 1 \end{bmatrix}\), \(\gamma = \begin{bmatrix} \sigma^2 \\ \rho \end{bmatrix}\), \(\sigma^2 > 0\), and \(\rho \in (-1, 1)\).
We can use numerical methods to find \(\hat\gamma\) , a maximizer of \(\ell^*(\gamma\mid y)\).
Example
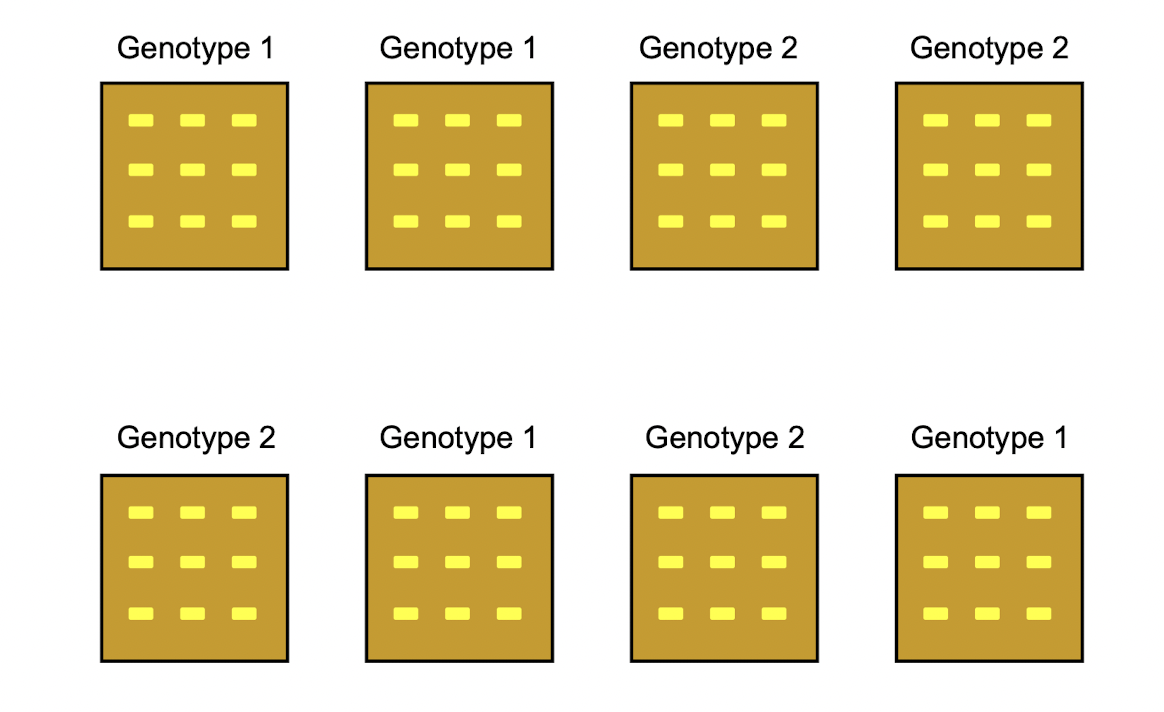
Researchers were interested in comparing the dry weight of maize seedlings from two different genotypes. For each genotype, nine seeds were planted in each of four trays. The eight trays in total were randomly positioned in a growth chamber. Three weeks after the emergence of the first seedling, emerged seedlings were harvested from each tray and individually weighed after drying to obtain one dry weight for each seedling. Although nine seeds were planted in each tray, fewer than nine seedlings emerged in many of the trays.
 |
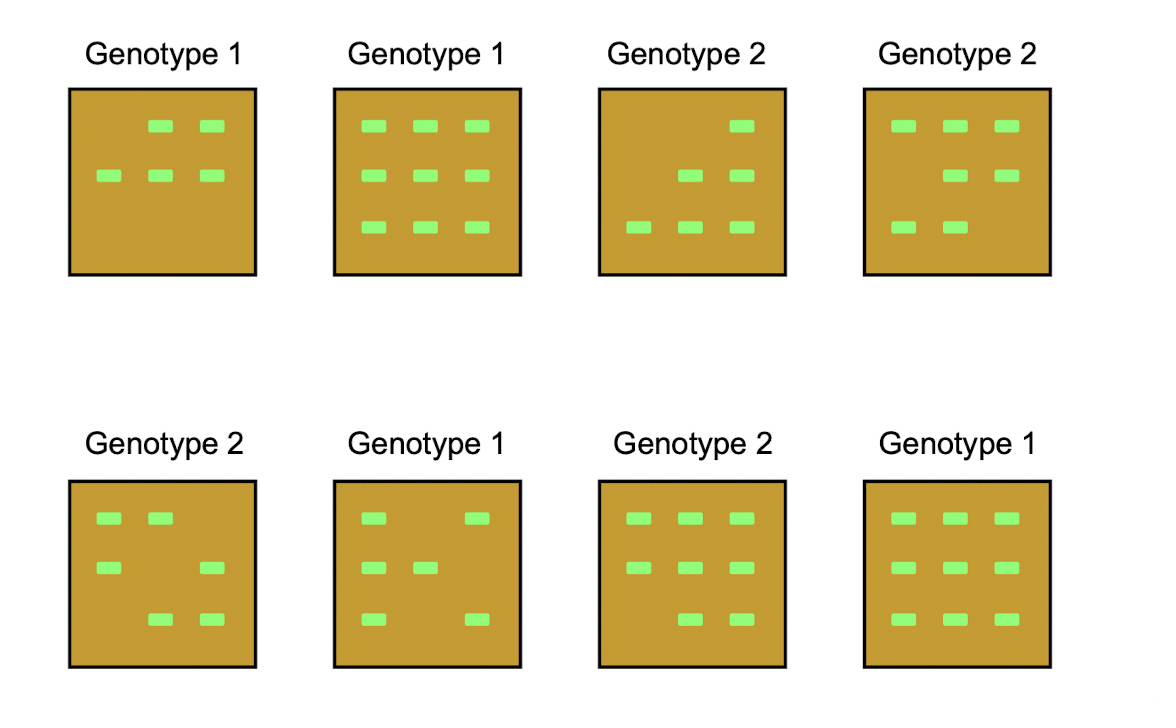 |
Let \(y_{ijk}\) be the dry weight of the \(k\)th seedling in the \(j\)th tray for genotype \(i\). Suppose \[ y_{ijk} = \mu_i + t_{ij} + e_{ijk} \] where \(\mu_1\) and \(\mu_2\) are unknown constants, \(t_{ij} \sim N(0, \sigma_t^2)\), \(e_{ijk} \sim N(0, \sigma_e^2)\), and all random terms are independent.
library(nlme)
lme(SeedlingWeight ̃Genotype,random= ̃1|Tray,method="ML", data=d)
Linear mixed-effects model fit by maximum likelihood
Data: d
Log-likelihood: -126.3709 # l(thetahat|y)
Fixed: SeedlingWeight ̃ Genotype
(Intercept) Genotype2
15.301832 -3.567017
Random effects:
Formula: ̃1 | Tray
(Intercept) Residual
StdDev: 2.932294 1.88247 # sigma_that, sigma_ehat
Number of Observations: 56
Number of Groups: 8
library(lme4)
lmer(SeedlingWeight ̃Genotype+(1|Tray),REML=F,data=d)
Linear mixed model fit by maximum likelihood [’lmerMod’]
Formula: SeedlingWeight ̃ Genotype + (1 | Tray)
Data: d
AIC BIC logLik deviance
260.7418 268.8432 -126.3709 252.7418
Random effects:
Groups Name Std.Dev.
Tray (Intercept) 2.932
Residual 1.882
Number of obs: 56, groups: Tray, 8
Fixed Effects:
(Intercept) Genotype2
15.302 -3.567