3 Prediction Using GAM
3.1 Bayesian Network
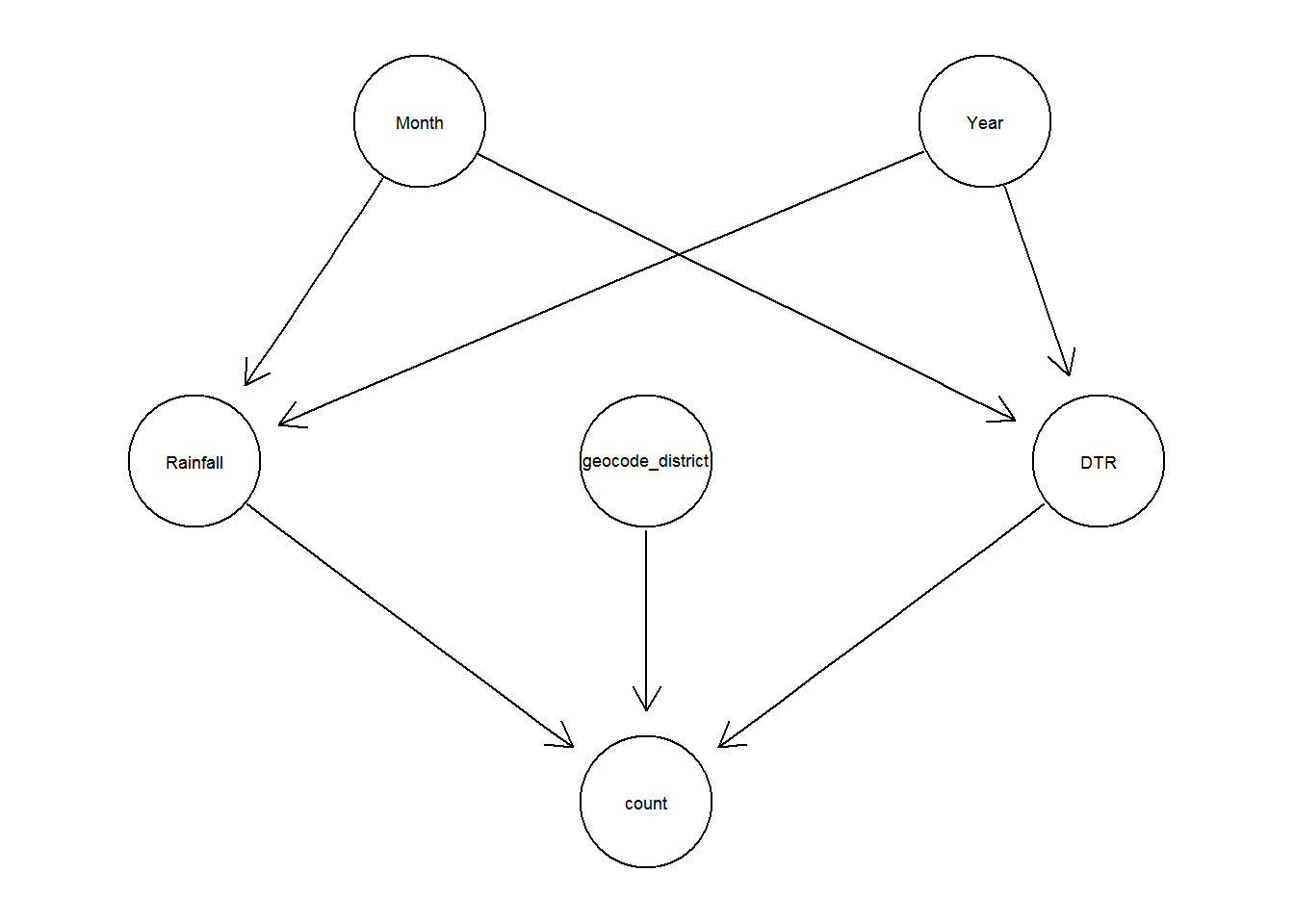
The accuracy of the prediction using Bayesian Network:
## ME RMSE MAE MPE MAPE
## Test set 9.386289 22.41716 13.33112 -26.19927 88.27519The in-sample error is
a <- sqrt(mean((trainCount-pred_train)^2))/sqrt(mean((trainCount)^2))
#for validation data 2013-2015
c <- sqrt(mean((testCount-pred_test)^2))/sqrt(mean((testCount)^2))The out-sample error for validation data 2013-2015 is
#for validation data 2013-2015
c <- sqrt(mean((testCount-pred_test)^2))/sqrt(mean((testCount)^2))The mean absolute percentage error:
mape <- function(y, yhat){
mean(abs((y - yhat)/y), na.rm=T) * 100
}
mpe <- function(y, yhat){
mean((y - yhat)/y, na.rm=T) * 100
}
mape(testCount, pred_test)## [1] 88.27519mpe(testCount, pred_test)## [1] -26.199273.2 Generalized Additive (Mixed) Models
3.2.1 Meterological Data
In this model, the association of meterologiocal variables i.e. DTR and averrage monthly rainfall is considered. I call this Dengue-Meteorological model.
The above summary in Appendix B.2 suggests that all the temperature and rain lag variables are important factors. Let’s visualize the additive model in Figure 3.1.
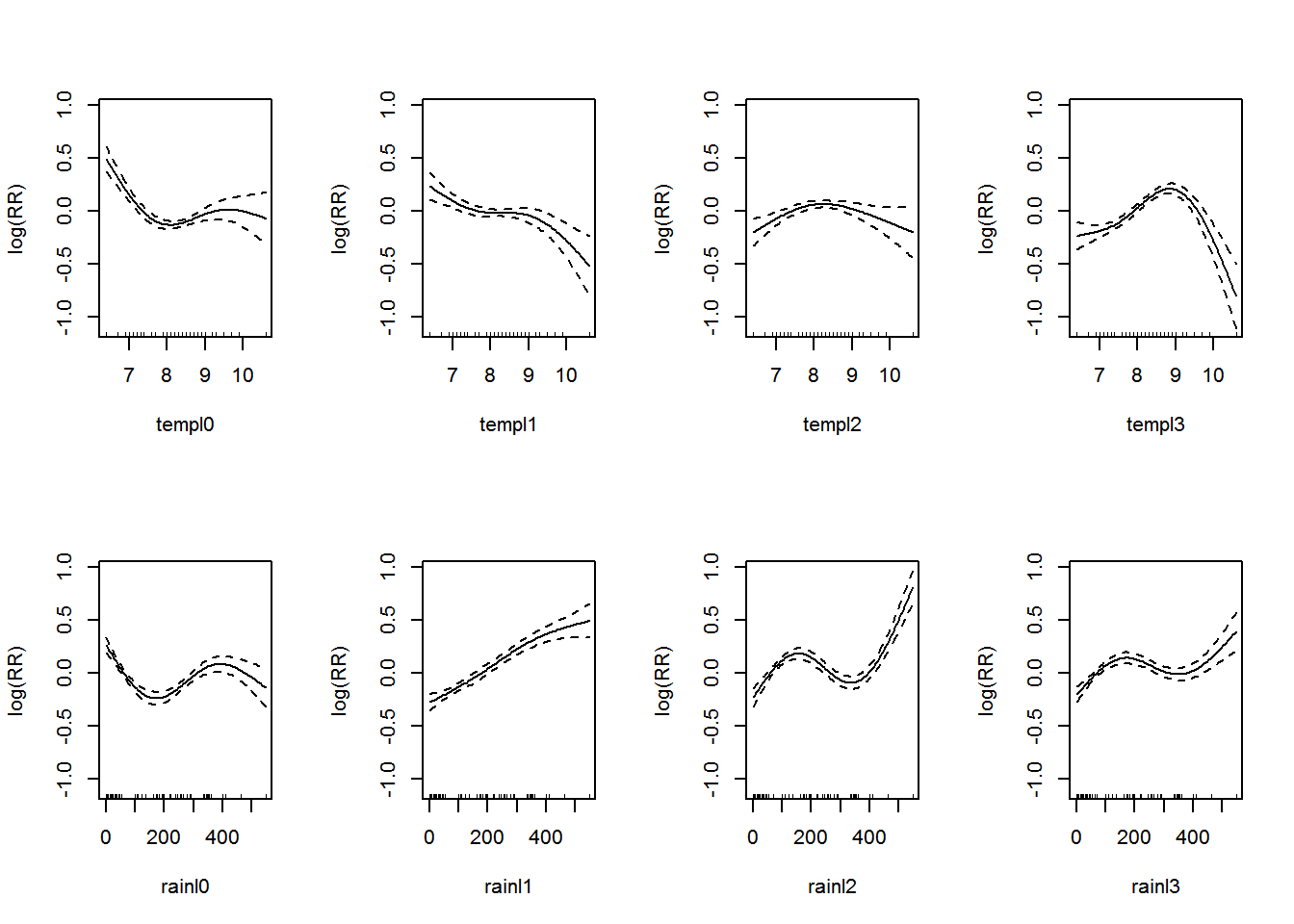
Figure 3.1: Association between the meteorological variables and dengue over lags of 0-3 months.. Solid lines represent relative risks (RR) of dengue cases and dottted lines depict the upper and lower limits of 95% confidence intervals.
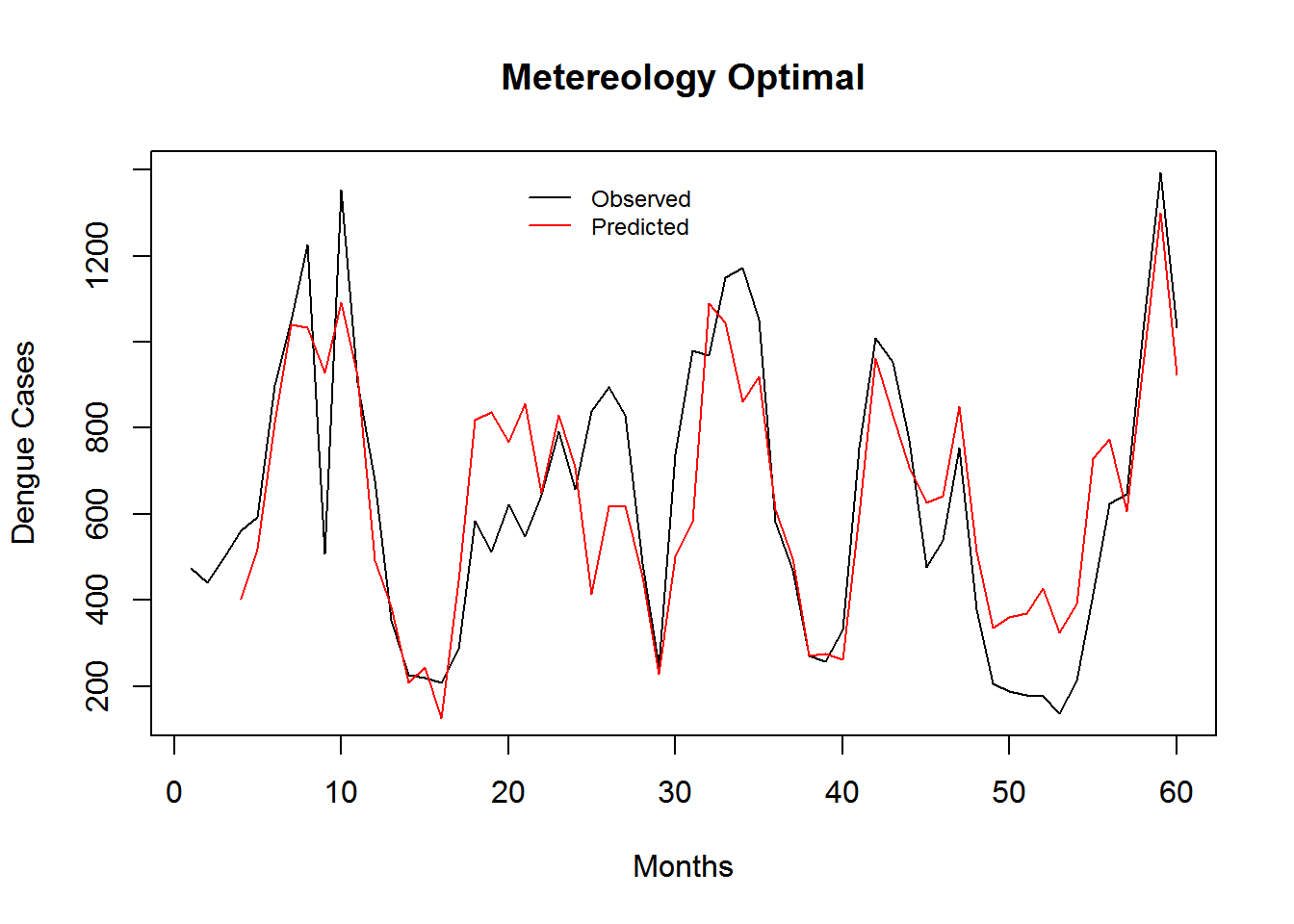
Figure 3.2: Monthly Observed and predicted dengue cases (2008-2012).
| Model Name | RMSE | SRMSE | R-sq.(adj) | Deviance Explained |
|---|---|---|---|---|
| Meteorology Model | 8.462372 | 0.5200771 | 0.2831722 | 0.3188542 |
3.2.2 Dengue Surveillance Data
In this model the association of past denge incidences is considered.
3.2.2.1 Short-term Lag Model
The summary of the model is shown in Appendix B.3. Let’s visualize the additive model in Figure 3.3.
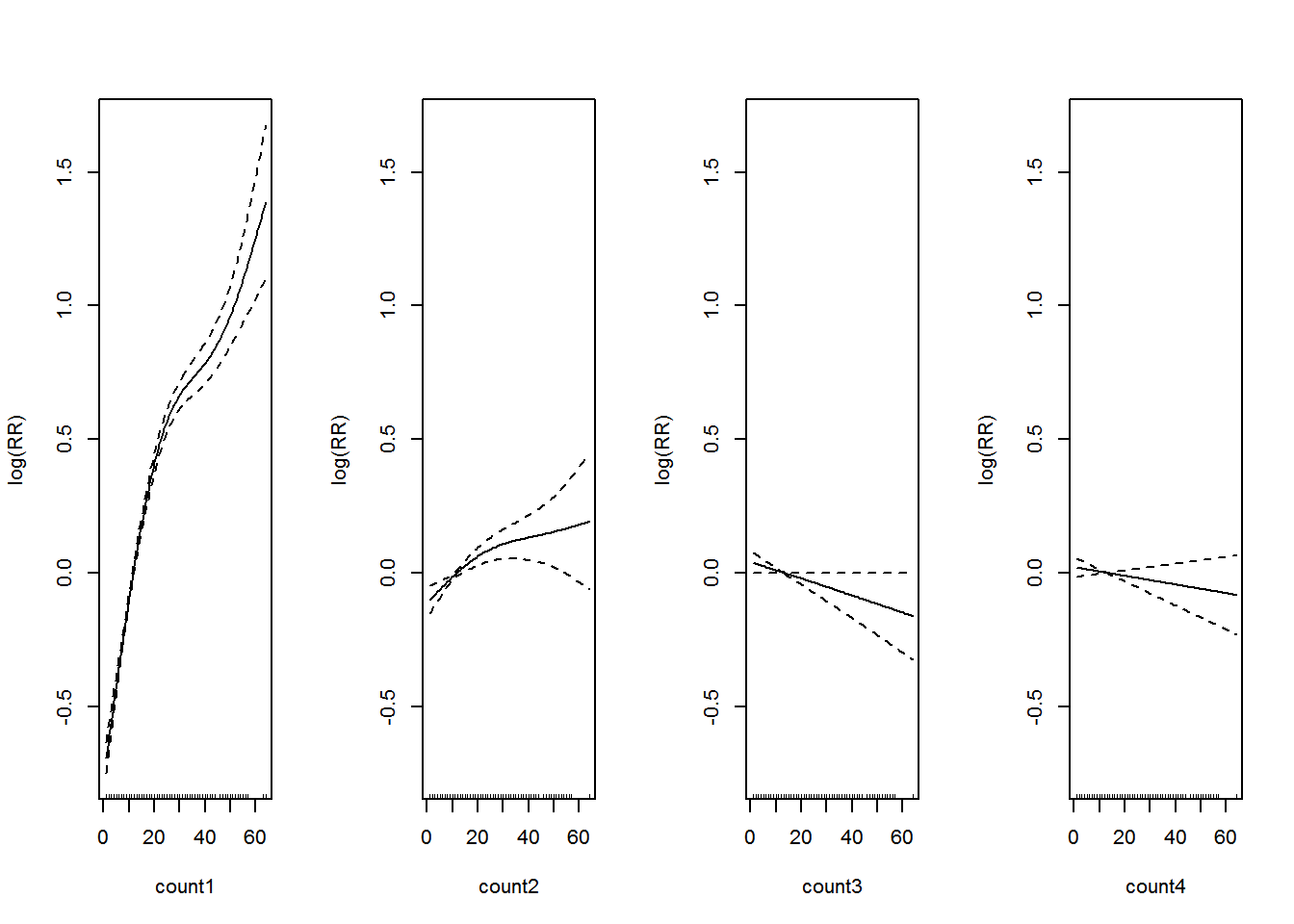
Figure 3.3: Association between past dengue count over lags of 1-4 months and the dengue outbreak.. Solid lines represent relative risks (RR) of dengue cases and dottted lines depict the upper and lower limits of 95% confidence intervals.
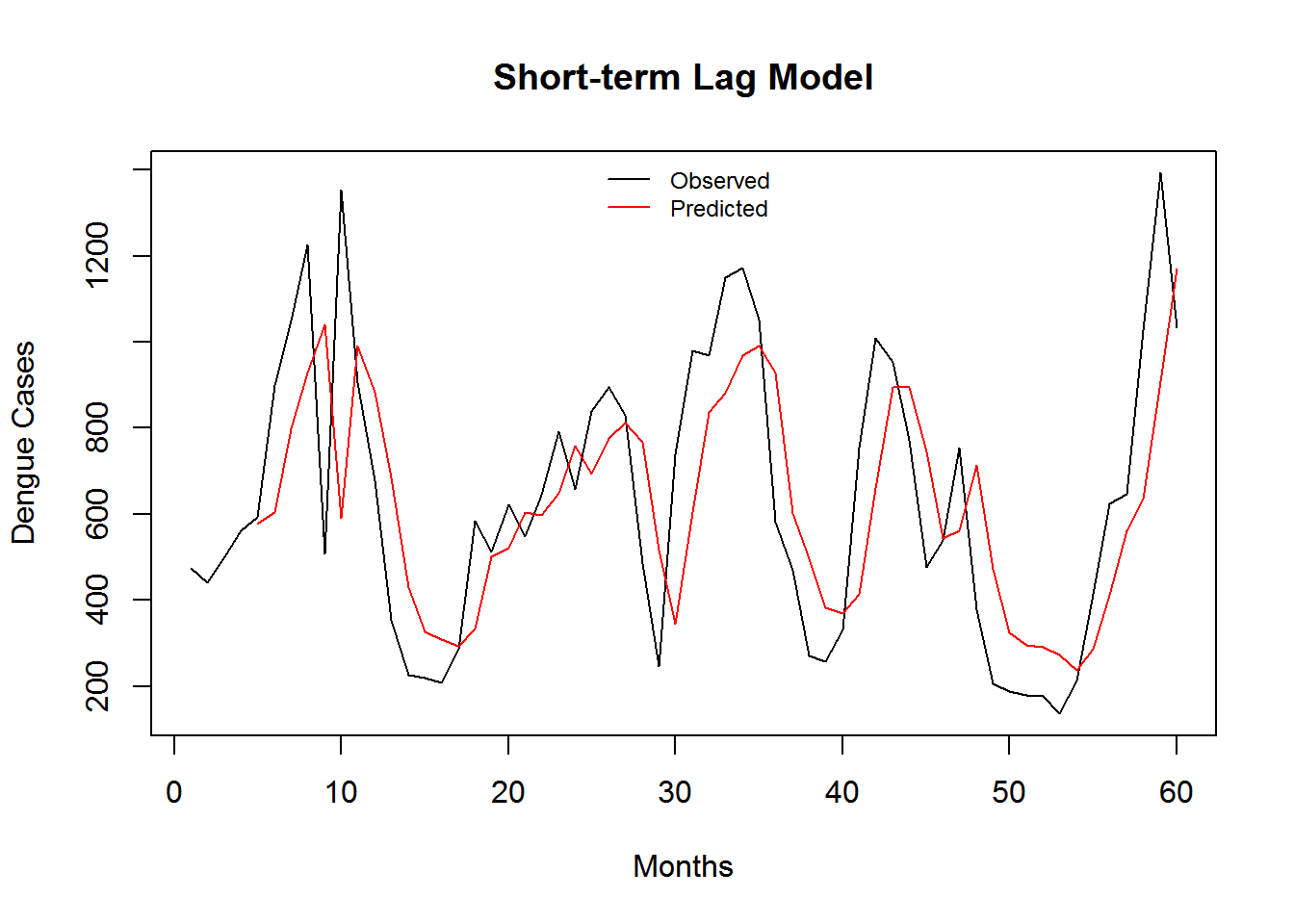
Figure 3.4: Monthly Observed and predicted dengue cases (2008-2012).
| Model Name | RMSE | SRMSE | R-sq.(adj) | Deviance Explained |
|---|---|---|---|---|
| Short-term Lag Model | 7.895124 | 0.4852154 | 0.3988267 | 0.4104035 |
3.2.2.2 Long-term Lag Model
## This is dlnm 2.2.6. For details: help(dlnm) and vignette('dlnmOverview').
## Important changes: see file.show(system.file('Changesince220',package='dlnm'))I show the simulated lag–response surfaces as relative risk in Figure 3.5.
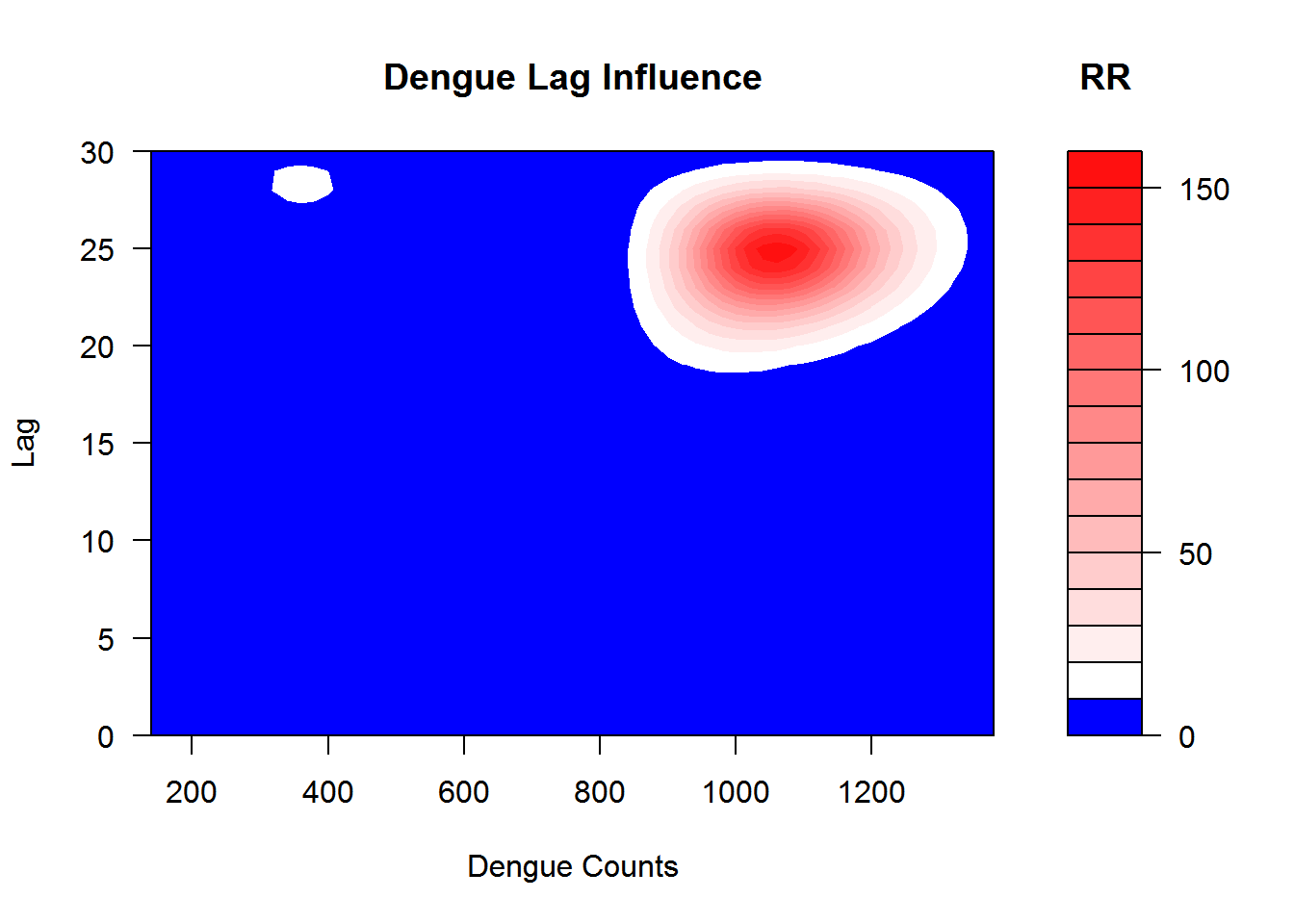
Figure 3.5: This shows the relation between the case intensity and dengue incidences at the lag months
The summary of the model is shown in Appendix B.5. Let’s visualize the additive model in Figure 3.6.
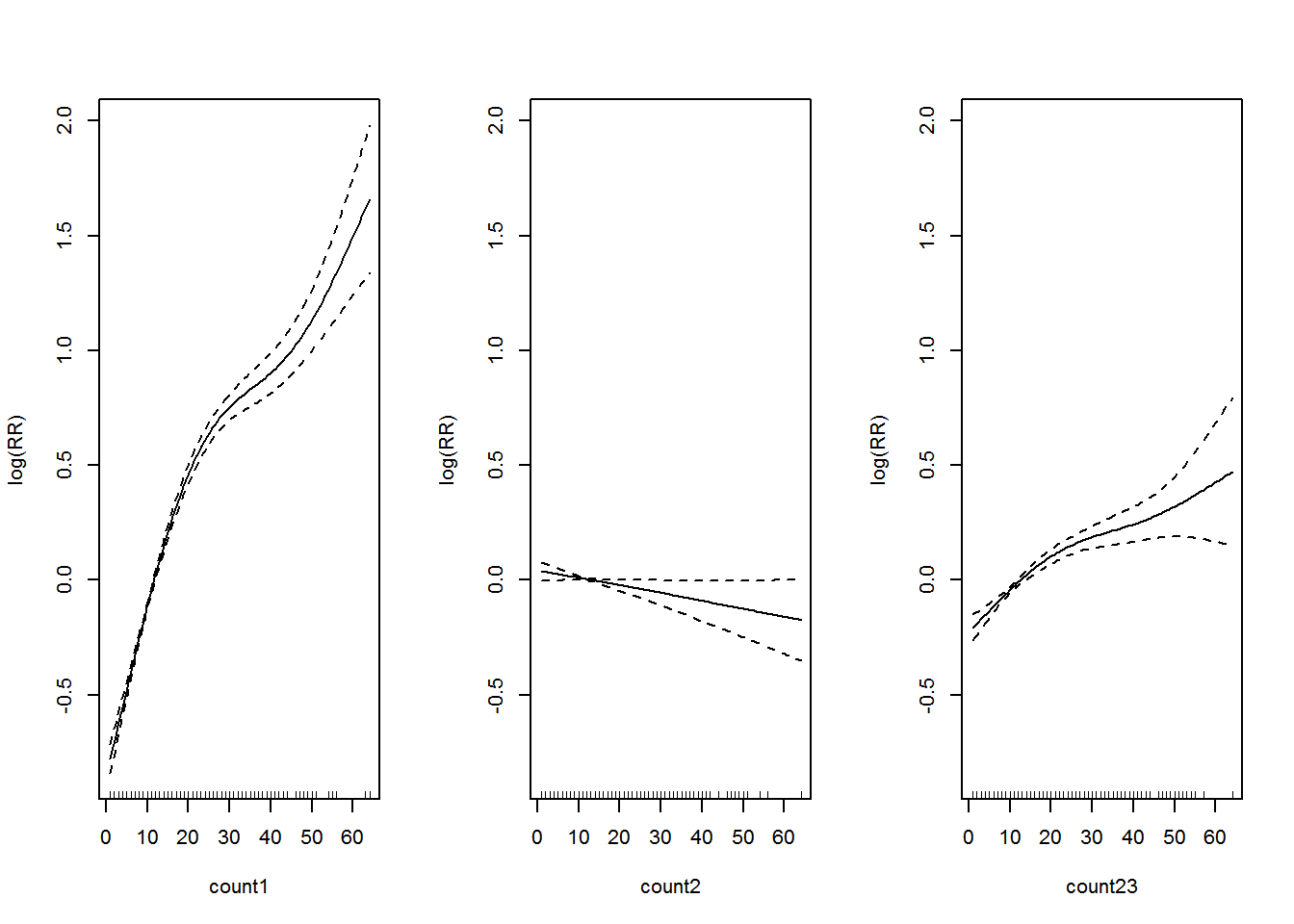
Figure 3.6: Association between past dengue count over optimal lags within 1-30 months and the dengue outbreak.. Solid lines represent relative risks (RR) of dengue cases and dottted lines depict the upper and lower limits of 95% confidence intervals.
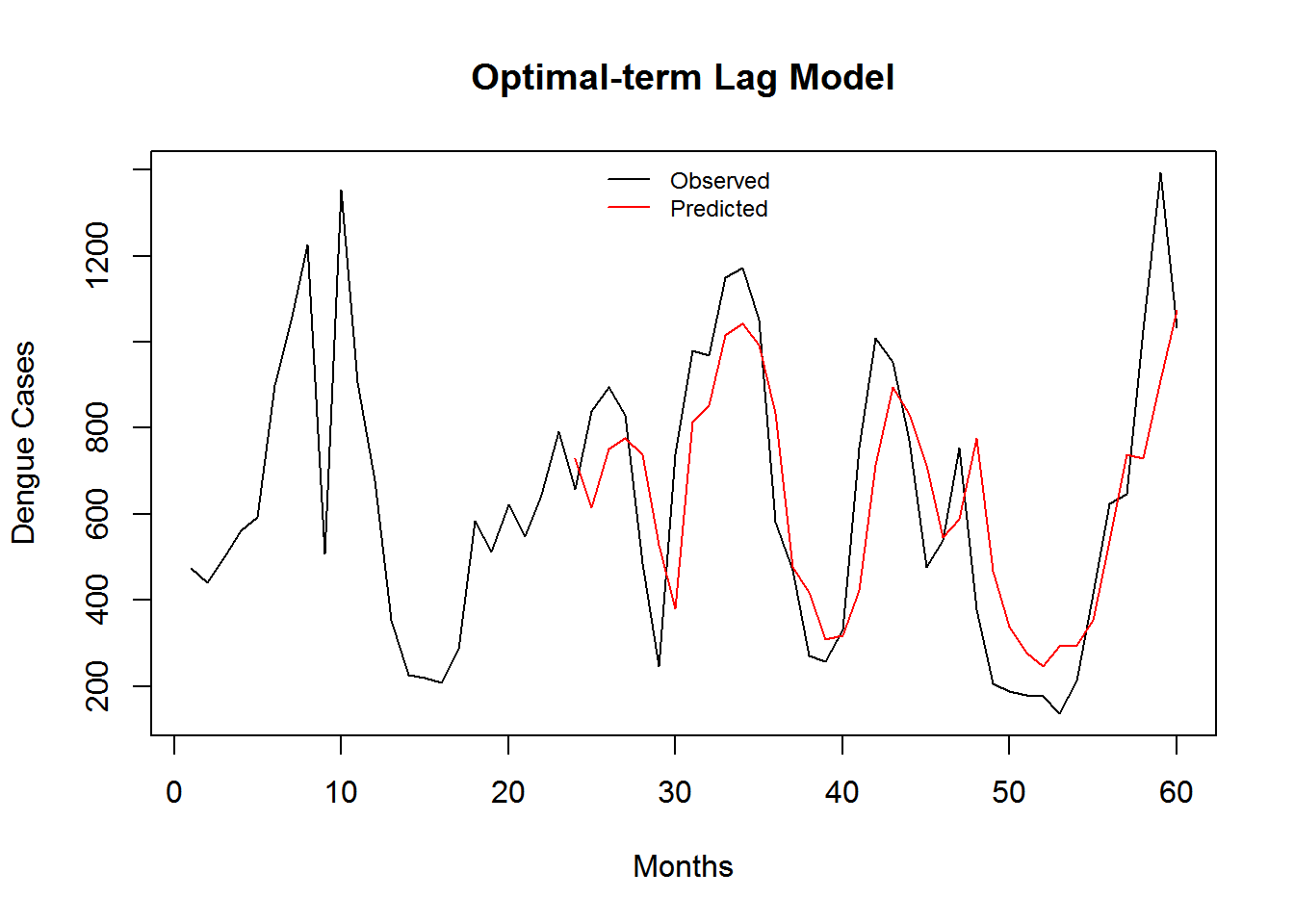
Figure 3.7: Monthly Observed and predicted dengue cases (2008-2012).
| Model Name | RMSE | SRMSE | R-sq.(adj) | Deviance Explained |
|---|---|---|---|---|
| Optimal-term Lag Model | 7.317273 | 0.4497021 | 0.4901485 | 0.488694 |
3.2.3 Meteorology and Optimal-term Lag Model.
The summary of the model is shown in Appendix B.6. Let’s visualize the additive model in Figure 3.8.
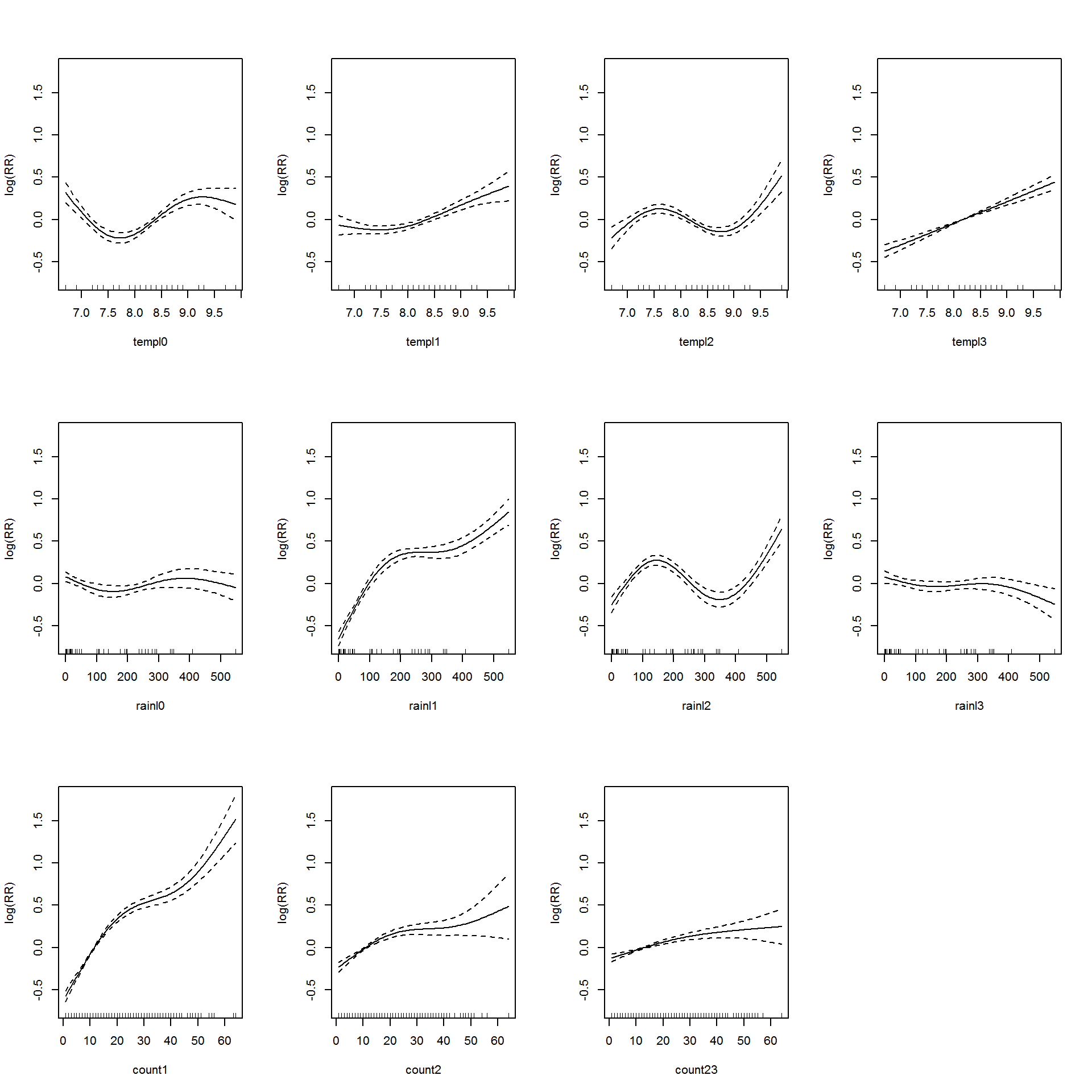
Figure 3.8: Association between the meteorological variables, past dengue count over optimal lags within 1-30 months and the dengue outbreak.. Solid lines represent relative risks (RR) of dengue cases and dottted lines depict the upper and lower limits of 95% confidence intervals.
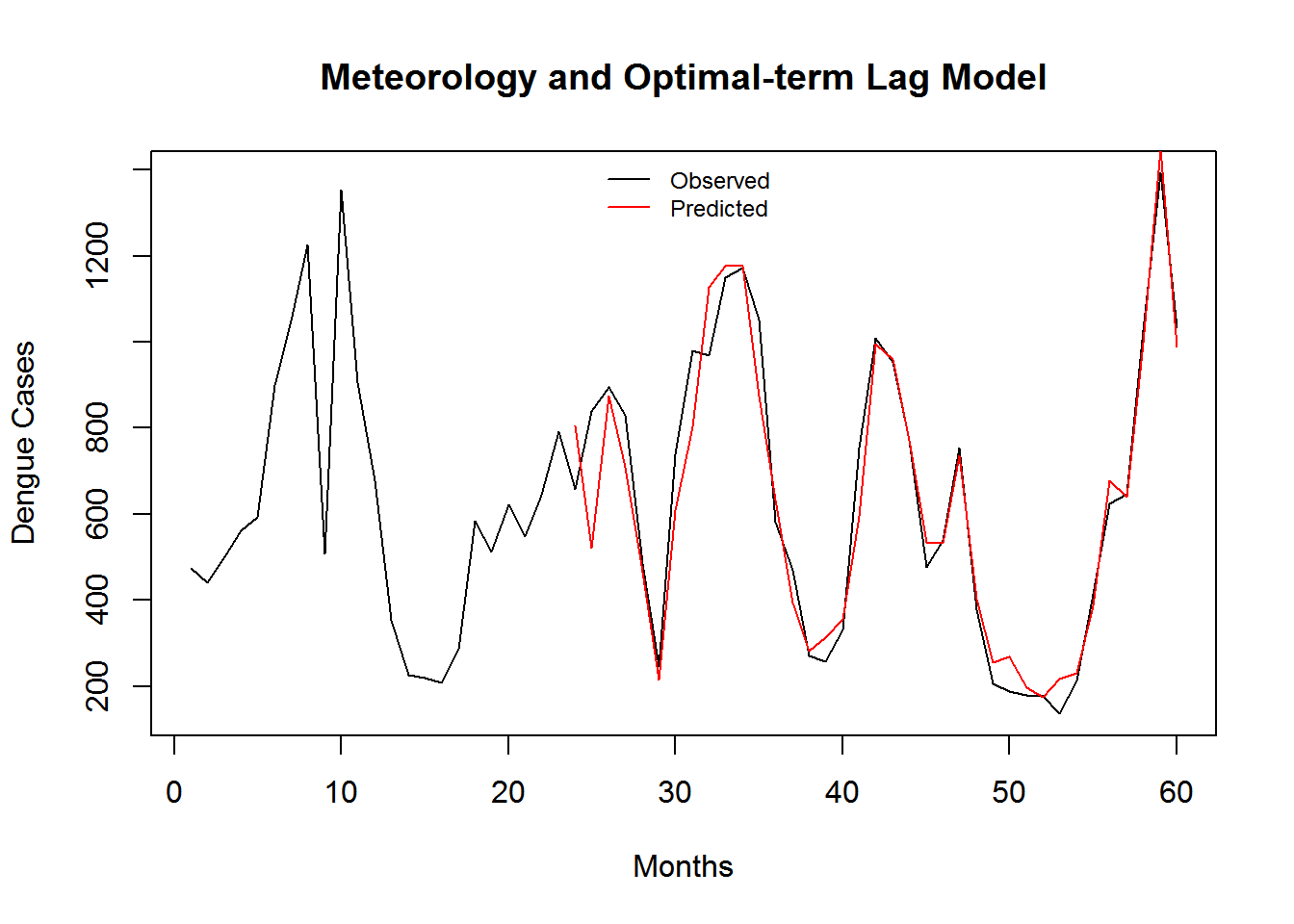
Figure 3.9: Monthly Observed and predicted dengue cases (2008-2012).
| Model Name | RMSE | SRMSE | R-sq.(adj) | Deviance Explained |
|---|---|---|---|---|
| Meteorology and Optimal-term Lag Model | 6.121665 | 0.3762228 | 0.6384466 | 0.6635623 |
3.2.4 Surrounding Dengue Data
I show the simulated lag–response surfaces for surrounding districts as relative risk in Figure 3.10.
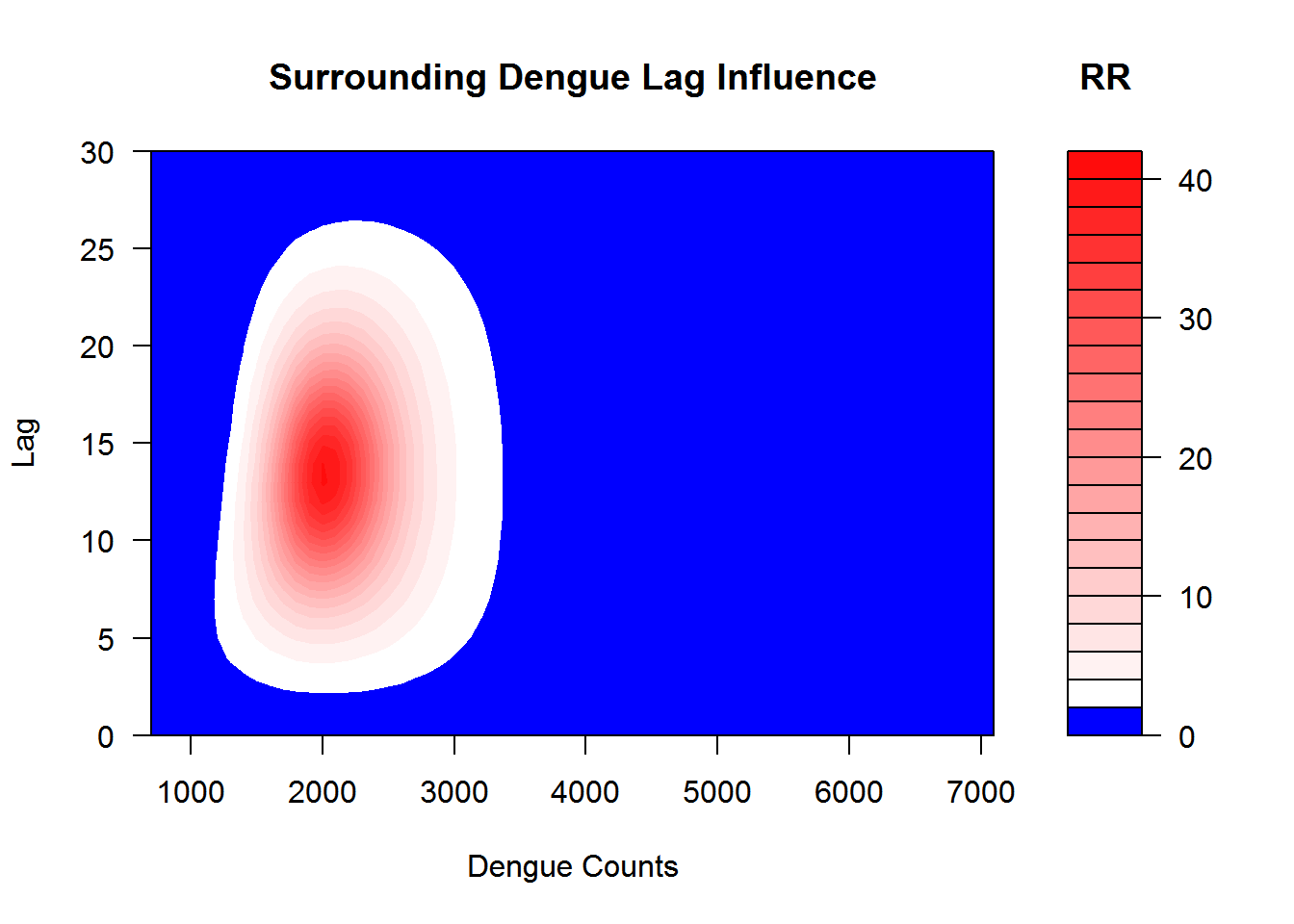
Figure 3.10: This shows the relation between the case intensity and dengue incidences in surrounding districts at the lag months
3.2.5 Meteorology, Optimal-term and Short-term Surrounding Lag Model
The summary of the model is shown in Appendix @ref(appDMDS_Short). Let’s visualize the additive model in Figure @ref(fig:DMDS_Short).
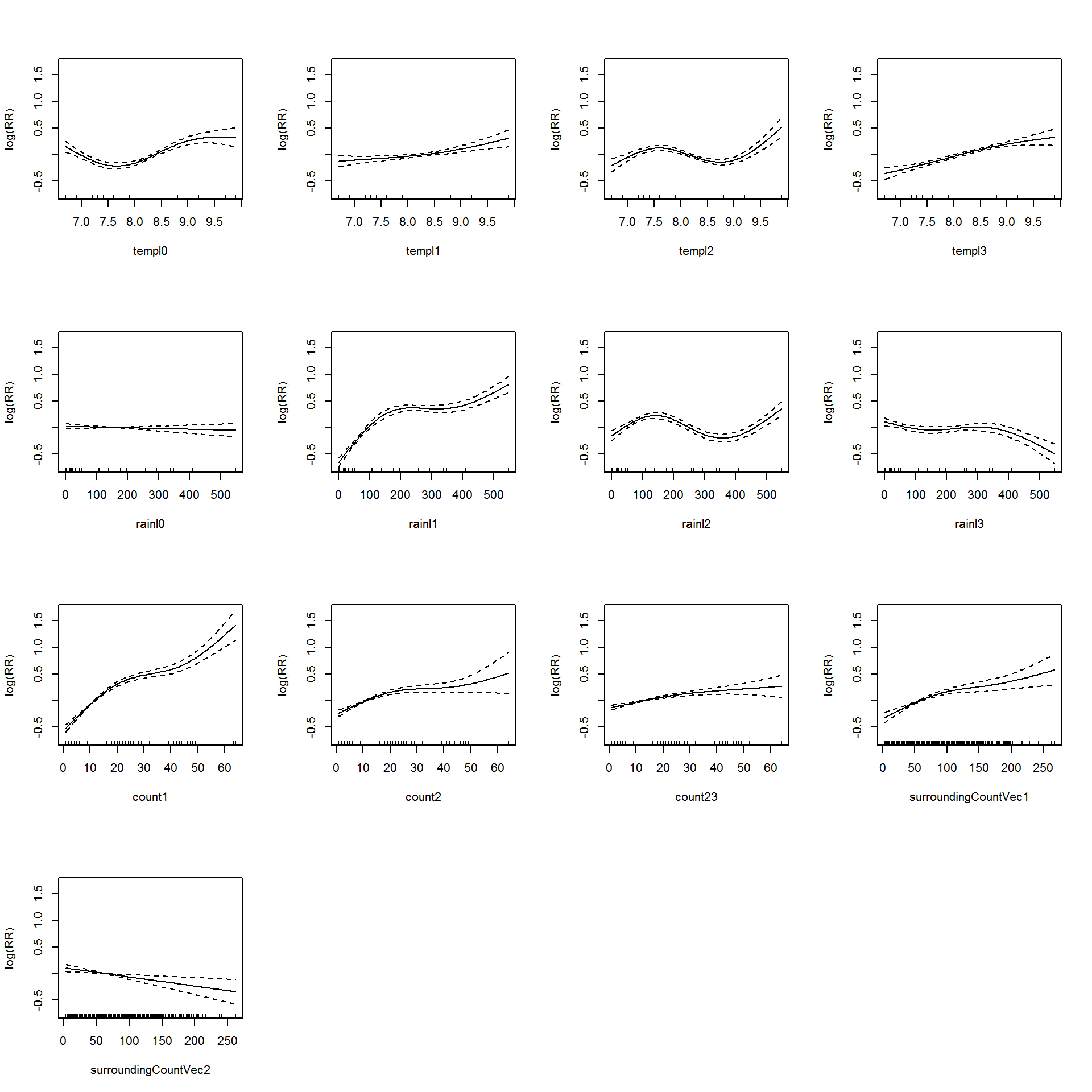
(#fig:DMDS_Short)Association between the meteorological variables, past dengue count over optimal lags within 1-30 months, surroinding district count over 0-3 months and the dengue outbreak.. Solid lines represent relative risks (RR) of dengue cases and dottted lines depict the upper and lower limits of 95% confidence intervals.
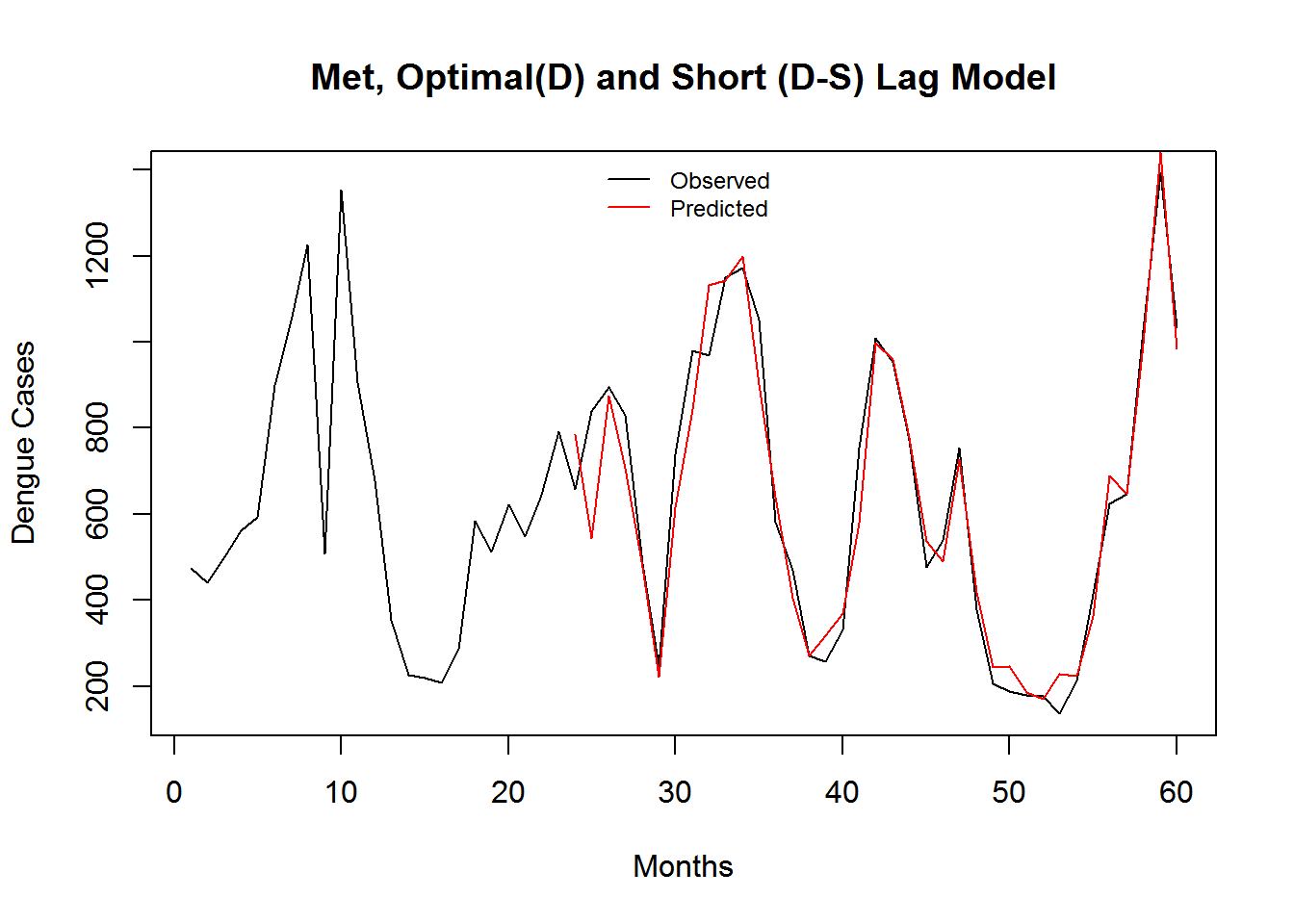
(#fig:DMDSShort_Pred)Monthly Observed and predicted dengue cases (2008-2012).
| Model Name | RMSE | SRMSE | R-sq.(adj) | Deviance Explained |
|---|---|---|---|---|
| Meteorology, Optimal(D) Short(D-S) Lag Model | 6.000072 | 0.36875 | 0.6521614 | 0.6726038 |
3.2.6 Meteorology, Optimal-term and Optimal-term Surrounding Lag Model
The summary of the model is shown in Appendix @ref(appDMDS_Optimal). Let’s visualize the additive model in Figure @ref(fig:DMDS_Optimal).

(#fig:DMDS_Optimal)Association between the meteorological variables, past dengue count over optimal lags within 1-30 months, surrounding district count over 0-30 months and the dengue outbreak.. Solid lines represent relative risks (RR) of dengue cases and dottted lines depict the upper and lower limits of 95% confidence intervals.
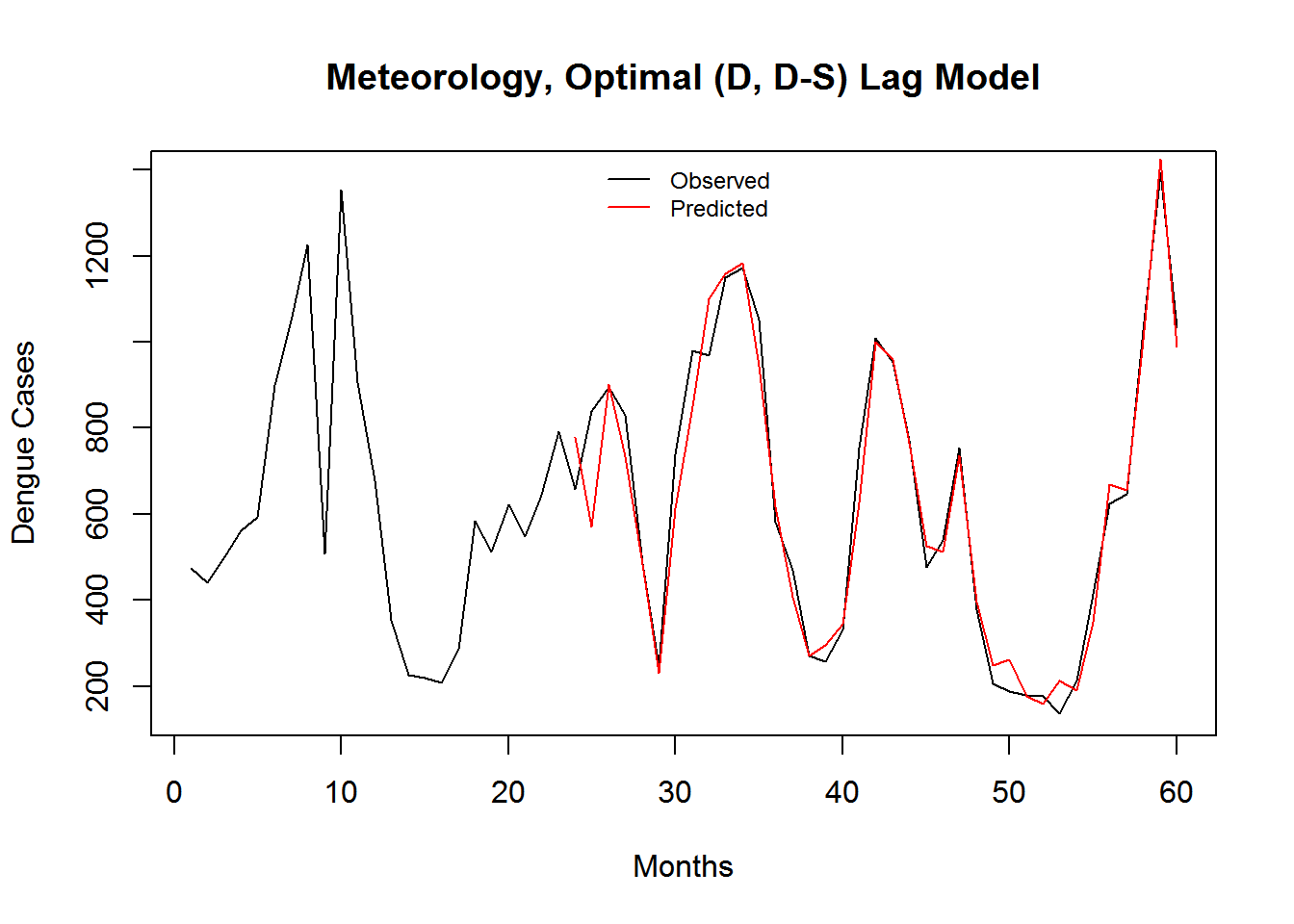
Figure 3.11: Monthly Observed and predicted dengue cases (2008-2012).
| Model Name | RMSE | SRMSE | R-sq.(adj) | Deviance Explained |
|---|---|---|---|---|
| Meteorology, Optimal (D, D-S) Lag Model | 5.940368 | 0.3650807 | 0.6581915 | 0.6809764 |
3.4 Predictive Performance Statistics
On the training dataset.
| Model Name | RMSE | SRMSE | R-sq.(adj) | Deviance Explained |
|---|---|---|---|---|
| Meteorology Model | 8.462372 | 0.5200771 | 0.2831722 | 0.3188542 |
| Short-term Lag Model | 7.895124 | 0.4852154 | 0.3988267 | 0.4104035 |
| Optimal-term Lag Model | 7.317273 | 0.4497021 | 0.4901485 | 0.4886940 |
| Meteorology and Optimal-term Lag Model | 6.121665 | 0.3762228 | 0.6384466 | 0.6635623 |
| Meteorology, Optimal(D) Short(D-S) Lag Model | 6.000072 | 0.3687500 | 0.6521614 | 0.6726038 |
| Meteorology, Optimal (D, D-S) Lag Model | 5.940368 | 0.3650807 | 0.6581915 | 0.6809764 |
| Social-economic data Included | 5.924147 | 0.3640839 | 0.6594216 | 0.7457561 |
3.5 Evaluation
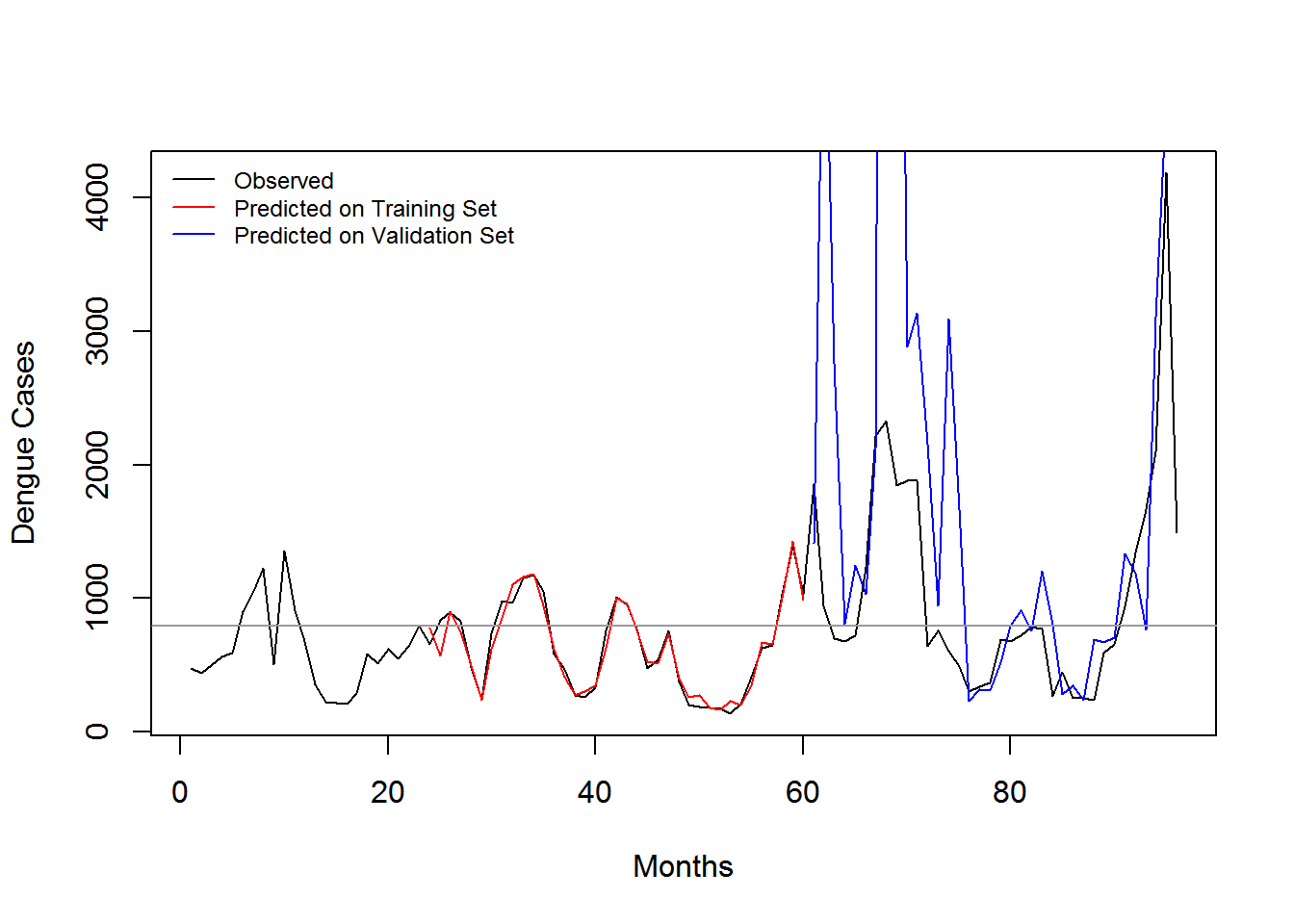
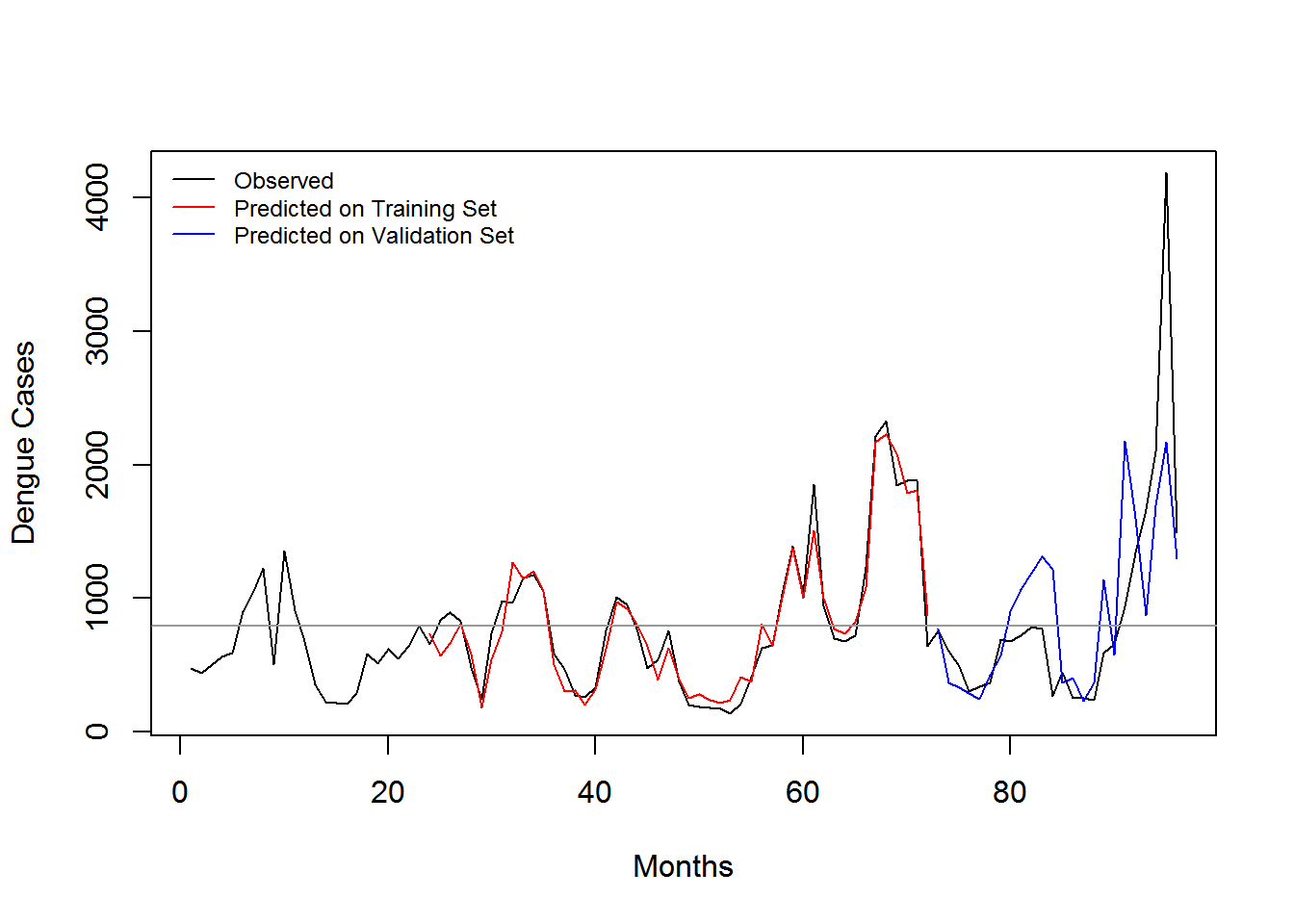
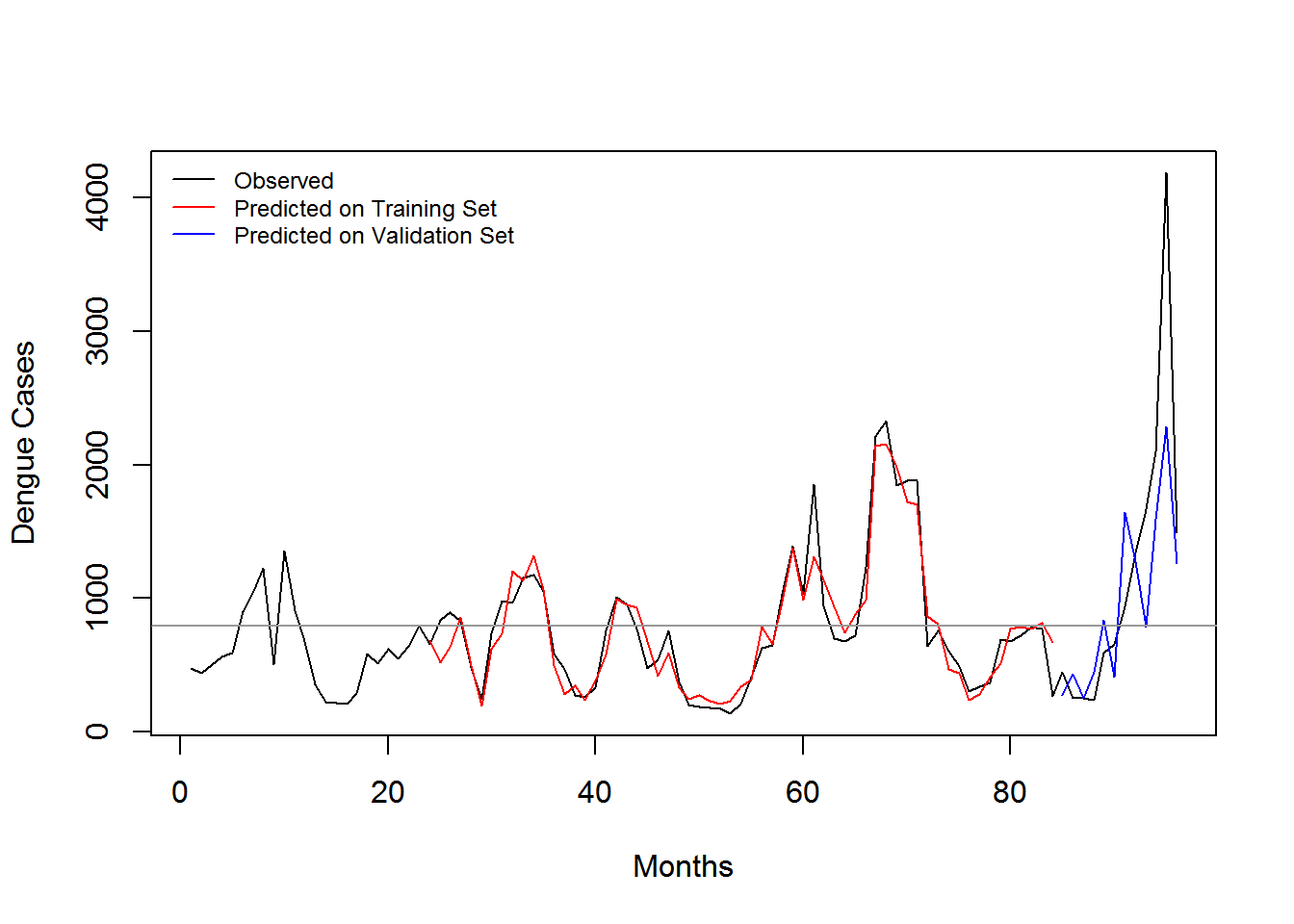
| Training Dataset | In-sample Error | Out-Sample (2013-2015) | Out-Sample (2014-2015) | Out-Sample (2013) | Out-Sample (2014) | Out-Sample (2015) |
|---|---|---|---|---|---|---|
| 2008-2012 | 0.3650807 | 303.282 | 410.7401247 | 21.66415 | 1.4334466 | 437.4985502 |
| 2008-2013 | 0.3843044 | NA | 0.5627701 | NA | 0.3202835 | 0.5354210 |
| 2008-2014 | 0.3940100 | NA | NA | NA | NA | 0.4888606 |
3.3 Social-Economic Data
The summary of the model is shown in Appendix B.9. Let’s visualize the additive model in Figure 3.12.
Figure 3.12: Association between the meteorological variables, past dengue count over optimal lags within 1-30 months, surrounding district count over 0-30 months, garbage data and the dengue outbreak.. Solid lines represent relative risks (RR) of dengue cases and dottted lines depict the upper and lower limits of 95% confidence intervals.
Figure 3.13: Monthly Observed and predicted dengue cases (2008-2012).