Chapter 3 Machine Learning in R
using the caret package
In Computer Labs 9B-11B we will use the caret R package (Kuhn et al. 2021) (short for Classification And REgression Training) to carry out machine learning tasks in RStudio.
The caret package offers a range of tools and models for classification and regression machine learning problems. In fact, it offers over 200 different machine learning models from which to choose. Don’t worry, we don’t expect you to use them all!
3.1 The caret package
We can download, install and load the caret package in RStudio as follows:
install.packages("caret")
library(caret)3.2 Predicting penguin species using machine learning
To illustrate an example application of the caret package, we will use the familiar penguins data set from the palmerpenguins R package (Horst, Hill, and Gorman 2020).
Suppose we would like to predict the species of penguins in the Palmer archipelago, based on their other characteristics - namely their bill_length_mm, bill_depth_mm, flipper_length_mm, body_mass_g and sex measurements (for this example we will ignore the other variables in the penguins data set).
Therefore, we have a multi-class classification problem, with the feature variables bill_length_mm, bill_depth_mm, flipper_length_mm, body_mass_g and sex, and the outcome variable species. Given we actually have recorded species observations already, our ML task can be categorised as a supervised learning task.
To begin, we load the palmerpenguins package (which should already be installed).
library(palmerpenguins)To make the following steps easier to follow, let’s create a data set containing only our feature and outcome variables (we will also remove missing values):
ml_penguins <- na.omit(penguins[, c(1,3:7)])3.2.1 Data Visualisation
As we know by now, it is usually a good idea to visualise our data before conducting any analyses. Since we should be quite familiar with the penguins data set, we won’t spend too long on this topic here.
However, it is worthwhile to note that the caret package offers several options for visualising data, via the featurePlot function. Here we use this function (with the argument plot = "pairs") to produce a scatter plot matrix of the different feature variables we are using, coloured by penguin species.
Note that the featurePlot function’s plot argument can take several different options, such as density, box, and scatter - you might like to try these out.
featurePlot(x = ml_penguins[, -1],
y = ml_penguins$species,
plot = "pairs",
auto.key = list(columns = 3))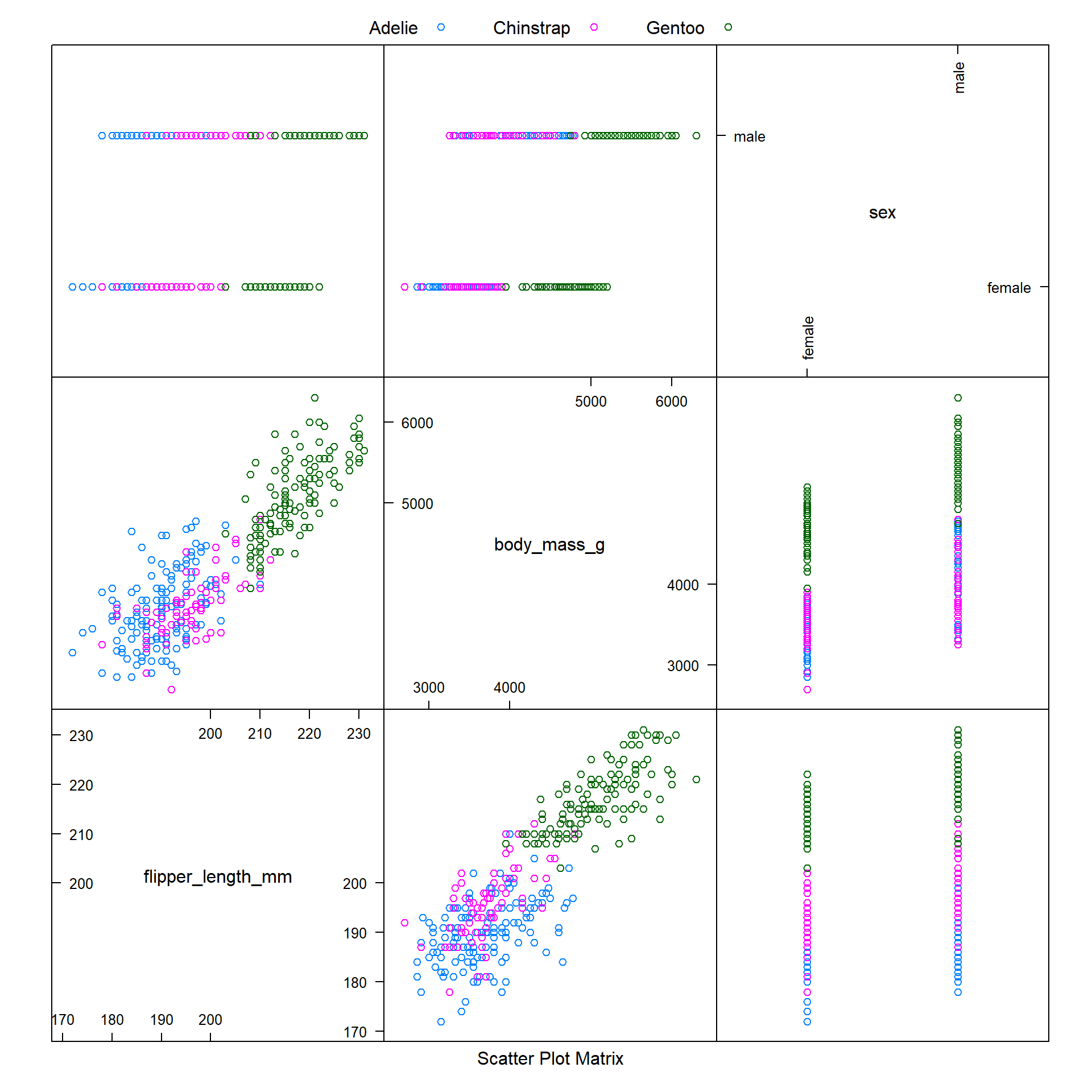
We observe that it is difficult to distinguish between Adelie and Chinstrap penguins when modelling body_mass_g against flipper_length_mm or bill_depth_mm. But hopefully our machine learning model will be able to use the data for these variables to make accurate predictions.
3.2.2 Pre-Processing
The caret package contains several tools for pre-processing, which makes our job easier.
3.2.2.1 Dummy Variables
One assumption made by the package is that all the feature variable data are numeric. Since our sex variable is categorical rather than numeric, we will have to convert it to a numeric variable before continuing.
We can use the dummyVars function from the caret package to reclassify the penguin sex recordings as ‘dummy variables’ (i.e. variables that take values 0 or 1, depending on whether they are true or not). We will call this adjusted data set dummy_penguins.
When converting feature variables via the dummayVars function, we need to follow a specific approach:
- First we assign the output of the
dummyVarsfunction to an object - Then we use that object, with the
predictfunction, and the original data (specified via thenewdataargument in thepredictfunction), to update our data - remember to also include the outcome variable when updating your data!
Let’s take a look at how we do this in R:
library(tibble)
dummy_penguins <- dummyVars(species ~ ., data = ml_penguins)
ml_penguins_updated <- as_tibble(predict(dummy_penguins, newdata = ml_penguins))
# remember to include the outcome variable too
ml_penguins_updated <- cbind(species = ml_penguins$species, ml_penguins_updated)
head(ml_penguins_updated)## species bill_length_mm bill_depth_mm flipper_length_mm body_mass_g sex.female
## 1 Adelie 39.1 18.7 181 3750 0
## 2 Adelie 39.5 17.4 186 3800 1
## 3 Adelie 40.3 18.0 195 3250 1
## 4 Adelie 36.7 19.3 193 3450 1
## 5 Adelie 39.3 20.6 190 3650 0
## 6 Adelie 38.9 17.8 181 3625 1
## sex.male
## 1 1
## 2 0
## 3 0
## 4 0
## 5 1
## 6 0Note: We use the as_tibble function from the tibble package to restructure our data following the introduction of the dummyVars dummy variables. This is mainly because we would like to include the species variable with the labels Adelie, Chinstrap and Gentoo, rather than the numbers 1,2 and 3.
Now, instead of sex taking the values of female or male, this variable has been replaced by the dummy variables sex.female and sex.male. Notice that in the first row, we have a value of 0 for sex.female and a value of 1 for sex.male - in other words, the data in the first row is for a male penguin.
3.2.2.2 Identifying samples exerting excessive influence
Before we begin training our machine learning model, we should also run some checks to ensure the quality of our data is high.
For instance, we should check our data to ensure that:
- Our data is balanced, with only a small number of unique values (if any) for each feature variable
- There are no samples that might have an excessive influence on the model
- We do not have any highly correlated feature variables\(^\dagger\)
\(^\dagger\)Sometimes, a machine learning model will benefit from using training data which includes several highly correlated feature variables. Often however, correlated feature variables can be problematic.
3.2.2.3 Zero and Near-Zero Variance Feature Variables
If any feature variables have zero or near-zero variance, this can cause problems when we subsequently split our data into training and validation data sets.
We can use the nearZeroVar function from the caret package to check a and b on our checklist.
One of the arguments of this function is saveMetrics, which can be specified as either saveMetrics = F or saveMetrics = T.
If we use saveMetrics = F, a vector of the positions of the feature variables with zero or near-zero variance will be produced.
If we use saveMetrics = T, a data frame with details about the variables will be produced.
Let’s consider both options, using our ml_penguins_updated data set.
saveMetrics = F option
nearZeroVar(ml_penguins_updated, saveMetrics = F)## integer(0)The output integer(0) means that none of the feature variables have been flagged as problematic, with respect to zero variance or near zero variance, which is encouraging. This means that none of the feature variables have only a single unique value.
saveMetrics = T option
nearZeroVar(ml_penguins_updated, saveMetrics = T)## freqRatio percentUnique zeroVar nzv
## species 1.226891 0.9009009 FALSE FALSE
## bill_length_mm 1.166667 48.9489489 FALSE FALSE
## bill_depth_mm 1.200000 23.7237237 FALSE FALSE
## flipper_length_mm 1.235294 16.2162162 FALSE FALSE
## body_mass_g 1.200000 27.9279279 FALSE FALSE
## sex.female 1.018182 0.6006006 FALSE FALSE
## sex.male 1.018182 0.6006006 FALSE FALSEHere, we can see that as identified previously, none of the variables have zero or near zero variance (as shown in columns 3 and 4 of the output).
The freqRatio column computes the frequency of the most prevalent value recorded for that variable, divided by the frequency of the second most prevalent value. If we check this column, we see that all feature variables have a freqRatio value close to 1. This is good news, and means that we don’t have an unbalanced data set where one value is being recorded significantly more frequently than other values.
Finally, if we check the percentUnique column, we see the number of unique values recorded for each variable, divided by the total number of samples, and expressed as a percentage. If we only have a few unique values (i.e. the feature variable has near-zero variance) then the percentUnique value will be small. Therefore, higher values are considered better, but it is worth noting that as our data set increases in size, this percentage will naturally decrease.
Based on these results, we can see that none of the variables show concerning characteristics.
All the variables have
freqRatiovalues close to 1.The
species,sex.maleandsex.femalevariables have lowpercentUniquevalues, but this is to be expected for these types of variables (if they were continuous numeric variables, then this could be cause for concern). In other words, categorical variables, e.g. dummy variables, often have lowpercentUniquevalues. This is normal and a lowpercentUniquevalue for a categorical feature variable is not by itself sufficient reason to remove the feature variable.
3.2.2.4 Cut-off Specifications
If we know beforehand that we have certain requirements for the freqRatio and percentUnique values, we can specify cut-off values using the arguments freqCut and uniqueCut respectively.
For example, if we considered feature variables with freqRatio scores higher than 1.23 and percentUnique scores lower than 20 to be exerting excessive influence, we could use the following code to filter out such feature variables:
nearZeroVar(ml_penguins_updated, saveMetrics = T, freqCut = 1.23, uniqueCut = 20)## freqRatio percentUnique zeroVar nzv
## species 1.226891 0.9009009 FALSE FALSE
## bill_length_mm 1.166667 48.9489489 FALSE FALSE
## bill_depth_mm 1.200000 23.7237237 FALSE FALSE
## flipper_length_mm 1.235294 16.2162162 FALSE TRUE
## body_mass_g 1.200000 27.9279279 FALSE FALSE
## sex.female 1.018182 0.6006006 FALSE FALSE
## sex.male 1.018182 0.6006006 FALSE FALSENotice how the output in the nzv column has changed compared to the initial output - now flipper_length_mm has an nzv value of TRUE, due to our arbitrary cut-off specifications.
3.2.2.4.1 Making conclusions based on cut-off values
In the event that a feature variable has both a high freqRatio value and a low percentUnique value, and both these values exceed the specified cut-offs, then it would be reasonable to remove this feature variable (assuming it is not a categorical variable).
If a feature variable has only one problematic value (e.g. a feature variable has a high freqRatio value that exceeds the specified cut-off, but also has a high percentUnique value which does not exceed the specified cut-off), then it is acceptable to retain this feature variable.
3.2.3 Data Splitting
Once we are happy with our data, we need to split it into training and validation data sets - we will call these ml_penguin_train and ml_penguin_validate respectively.
We can use the createDataPartition function from the caret package to intelligently split the data into these two sets.
One benefit of using this function to split our data compared to simply using the sample function is that if our outcome variable is a factor (like species!) the random sampling employed by the createDataPartition function will occur within each class.
In other words, if we have a data set comprised roughly 50% Adelie penguin data, 20% Chinstrap data and 30% Gentoo data, the createDataPartition sampling will preserve this overall class distribution of 50/20/30.
Let’s take a look at how to use this function in R:
set.seed(16505050)
train_index <- createDataPartition(ml_penguins_filtered$species,
p = .8, # here p designates the split - 80/20
list = FALSE,
times = 1) # times specifies how many splits to performHere we have split the training/validation data 80/20, via the argument p = 0.8. If we now take a quick look at our new object, we observe that:
head(train_index, n = 6)## Resample1
## [1,] 1
## [2,] 3
## [3,] 4
## [4,] 6
## [5,] 8
## [6,] 10Note that the observations 1, 3, 4, 6, 8 and 10 will now be assigned to the ml_penguin_train training data, while observations 2, 5 and 9 will be assigned to the ml_penguin_validate validation data.
To carry out these assignments using our train_index object, we can use the following code:
ml_penguin_train <- ml_penguins_filtered[train_index, ]
ml_penguin_validate <- ml_penguins_filtered[-train_index, ]We are now ready to train our model!
In the following section, we introduce a selection of machine learning models, which we will apply in Computer Labs 10B and 11B.
References
“Evil Carrot” by Brettf is licensed under CC BY 2.0↩︎
