Ex. 1
(a) Using “scan”
- Use
scan()to read every single cell in the baseball-salaries-2016.txt file, need to setsep="\t"to avoid a 2-gram term from being separated into two words. - Use
data.frame()to create a data.frame (class) by the input vector. - Then use
write.csv()andwrite.table()to export the file.
(b) Draw boxplots
- salary =
Salary ($M) - winning percentage (WPCT) = W/(W+L)
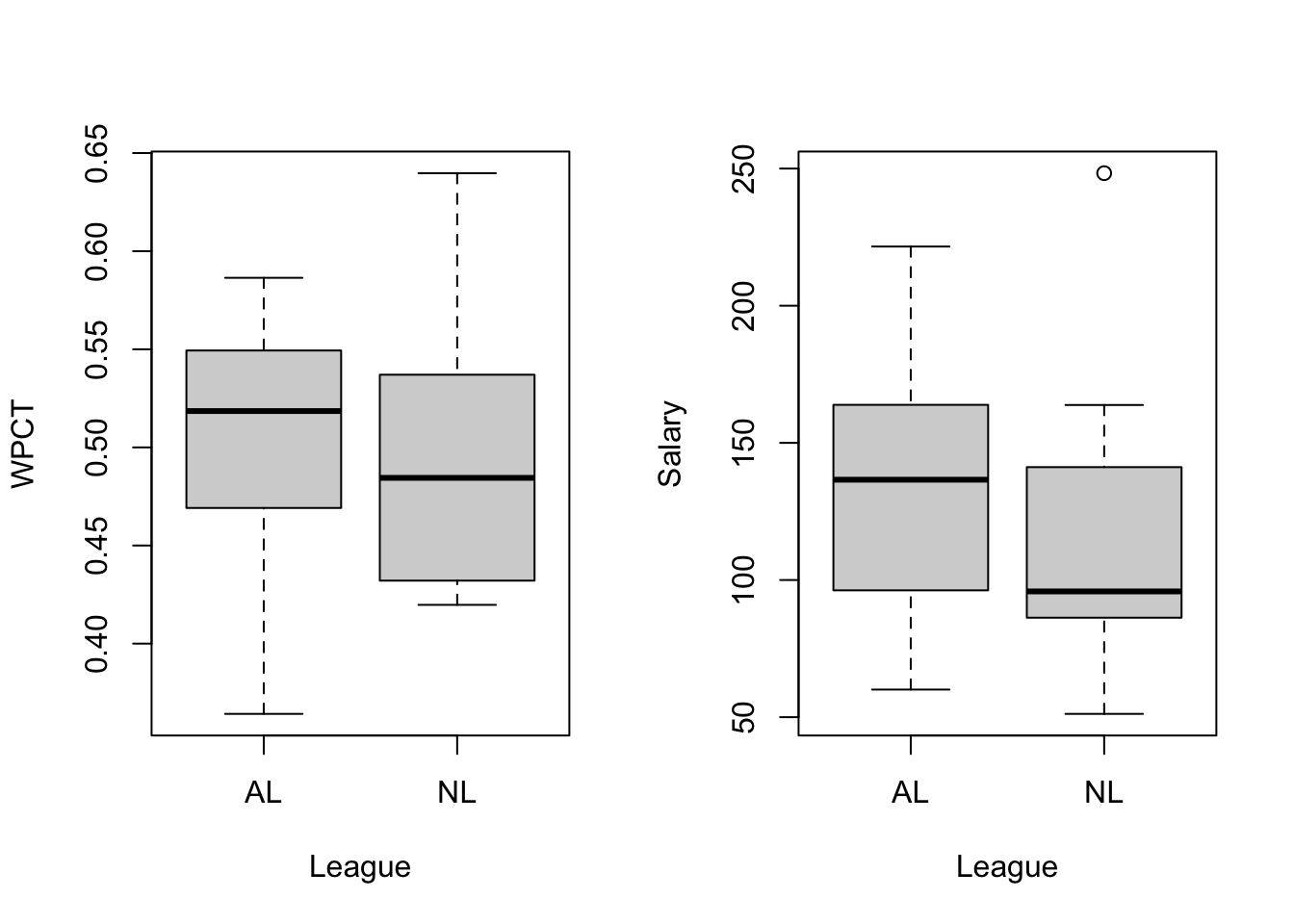
(c) Check correlation
- Using
cor.test()to check whether the positive relations between salary and winning percentage. - The Pearson’s correlation between salary and winning percentage is 0.617, p<.001, so we can reject the null hypothesis, i.e. the correlation between WPCT and salary is significant.
##
## Pearson's product-moment correlation
##
## data: sala and wpct
## t = 4.1436, df = 28, p-value = 0.0002856
## alternative hypothesis: true correlation is not equal to 0
## 95 percent confidence interval:
## 0.3294329 0.7992681
## sample estimates:
## cor
## 0.6165296(d) Scatter plot
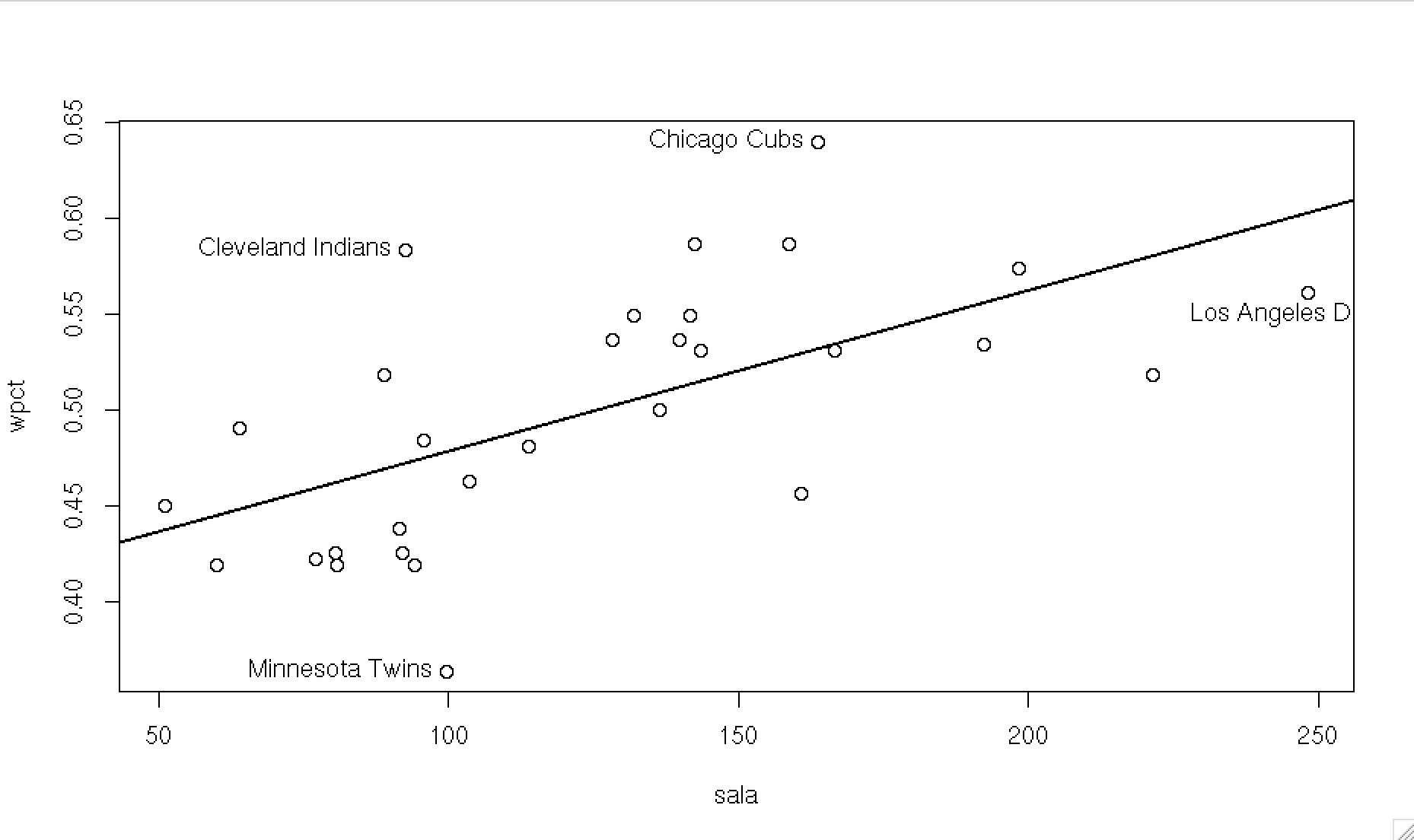
Ex. 2
(a) Length of a vector
The sample is shown below. When we shorten a vector (set a random seed) from length 20 to 15, only the first 15 numbers would be present. When we lengthen it to 18, three (18-15=3) NAs would be added.
- the results of
rnorm(20).
## [1] -1.21 0.28 1.08 -2.35 0.43 0.51 -0.57 -0.55 -0.56 -0.89
## [11] -0.48 -1.00 -0.78 0.06 0.96 -0.11 -0.51 -0.91 -0.84 2.42- the results of
length(x) = 15
## [1] -1.21 0.28 1.08 -2.35 0.43 0.51 -0.57 -0.55 -0.56 -0.89
## [11] -0.48 -1.00 -0.78 0.06 0.96- the results of
length(x) = 18
## [1] -1.21 0.28 1.08 -2.35 0.43 0.51 -0.57 -0.55 -0.56 -0.89
## [11] -0.48 -1.00 -0.78 0.06 0.96 NA NA NA(b) Operations of a vector
- Change the \(x<0\) part of
x[1:20]to NA. The results is shown below.
## [1] NA 0.28 1.08 NA 0.43 0.51 NA NA NA NA
## [11] NA NA NA 0.06 0.96 NA NA NA NA 2.42- computing the the mean, median, and variance of non-NA entries in
x[1:18]from (a).
## [1] "mean: 0.55"## [1] "median: 0.47"## [1] "var: 0.16"(c) Change NA to 0
- using
is.na()to identify the NAs, then replaced them by 0s.
## [1] 0.00 0.28 1.08 0.00 0.43 0.51 0.00 0.00 0.00 0.00
## [11] 0.00 0.00 0.00 0.06 0.96 0.00 0.00 0.00 0.00 2.42Ex. 3
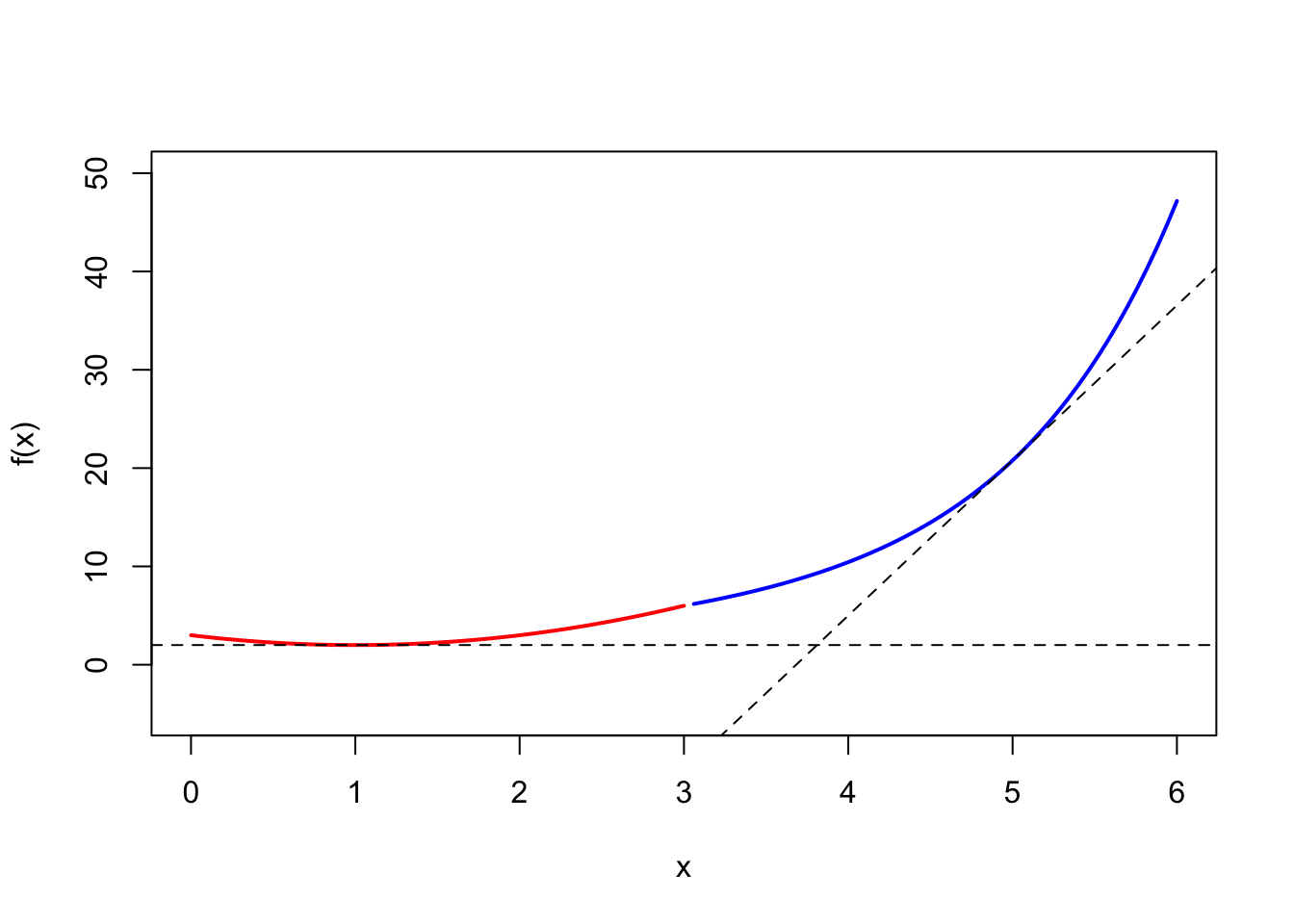
(a) Graph
The graph is shown as 3.(b) below.
(b) Tangent line
Find the slope and intercept of the tangent line of \(f(x)=x+2e^{(x-3)}+1\) at \(x=5\), then we can use abline() function to draw the tangent line. (Note. in the R abline() function, the intercept is denoted as a, and the slope is denoted as b.)
- Slope. \(f'(5)=1+2e^{(5-3)}=1+2e^2\)
- Point. \((x,y)=(5, f(5))=(5, 2e^2+6)\) ( Note. \(x=5\) substituted back into the original equation \(f(x)\))
- Intercept. based on \(y=bx+a\), and \(b=(1+2e^2)\) is known. (follow the parameters of
abline().)
\(2e^2+6=5(1+2e^2)+a\), find \(a=-8e^2+1\) .
# calculus
expression((x^2-2*x+3), 'x') |> D('x')## 2 * x - 2expression((x+2*exp(x-3)+1), 'x') |> D('x')## 1 + 2 * exp(x - 3)Ex. 4
(a) cat() and date()
The codes and outputs are shown below, combine the labels and system time by cat() function.
cat(
labels = "Today’s date is: ",
paste( substr(date(),1,10), substr(date(),21,24)),
fill = TRUE
)## Today’s date is: Tue Mar 22 2022cat(
labels = "The time now is: ",
paste( substr(date(),12,19)),
fill = TRUE
)## The time now is: 23:30:47(b) Explain the outputs of time.
- With the first 2 lines, we assign the system date to the object
today(with a class “Date”), then display it as day-month-year format, and the month%bis an abbreviated month in English. - With the 3rd line, we create a sequence that has 10 days, begins
today, and generates every other week. - In the 4th line, we extract the weekday name from the object
today. In the 5th line, we check every single month’s name from the sequencetenweekswith 10 values. - In the last line, we check the leap seconds (潤秒) before, then display them as the date format.
(today <- Sys.Date())
format(today, "%d %b %Y")
(tenweeks <- seq(today, length.out=10, by="1 week"))
weekdays(today)
months(tenweeks)
as.Date(.leap.seconds)Ex. 5
(a) Mid-Square Method
- By the requirements, the initial values is a 6 digits number, then square it.
- How to make sure the 12-digits (square of 6-digits) at the middle of number (including 0, ranging over [0, 999999])? We have introduced the padding method from the
stringrpackage. - The comment of the
gen_5a()function. (1) Create a null vectorvec. (2) In the for loop, we consider the input or iterate value may not be 6-digits, so we convert it to a character class first, then usestr_pad()function to impute 0 at the left side until 12 digits. (3) Usestr_sub()to extract the 6 digits from the 4th number to the 9th. (4) Convert the 6-digit number back into the numeric class. (5) Save the numbers in the vectorvecwith theith place. - Goodness-of-fit test. We generated 10000 random numbers from the
gen_5a()function. The K-S test showed D = 1, p-value < .001, so we reject the null hypothesis, i.e. the random numbers do not follow the uniform distribution.
## Warning in ks.test(res_5a, y = "punif"): ties should not be present for the
## Kolmogorov-Smirnov test##
## One-sample Kolmogorov-Smirnov test
##
## data: res_5a
## D = 1, p-value < 2.2e-16
## alternative hypothesis: two-sided(b) Vax’s generator
- The comment of the
gen_5b()function. (1) Create a null vectorvec. (2) In the for loop, implement a Vax’s generator(69069*x) %% (2^32). (3) Save the numbers in the vectorvecwith theith place. - Goodness-of-fit test. We generated 10000 random numbers from the
gen_5b()function. For \(\chi^2\) test, we separated the 10000 numbers to 10 groups by the intervals of 1/10 of range(max(res_5b) - min(res_5b))/k. Then check the K-S test D = 1, p < .001, we can reject the null hypothesis, i.e. the random numbers do not follow the uniform distribution. However, the \(\chi^2\) test showed \(\chi^2=6.9936\) with df=9, p=0.6378, we can not reject the null hypothesis, i.e. there are no significant difference between the random numbers and uniform distribution. - K-S test and \(\chi^2\) test have different results. But K-S test considers each value, may be better than \(\chi^2\) test.
##
## One-sample Kolmogorov-Smirnov test
##
## data: res_5b
## D = 1, p-value < 2.2e-16
## alternative hypothesis: two-sided##
## Chi-squared test for given probabilities
##
## data: chi_5b
## X-squared = 6.9936, df = 9, p-value = 0.6378(c) Multiplicative congruential generators
- The comment of the
gen_5c()function. (1) Create a null vectorvec. (2) In the for loop, implement a combined generators,(171*x) %% 30269,(172*y) %% 30307,(170*z) %% 30323, and(x/30269 + y/30307 + z/30323) %% 1. (3) Save the numbers in the vectorvecwith theith place. - Goodness-of-fit test. We generated 10000 random numbers from the
gen_5c()function. Check the K-S test D = 0.010053, p = 0.2643, we can not reject the null hypothesis, i.e. the random numbers do follow the uniform distribution. - Comparison of (a), (b) and (c). We can find the generator of 5(a) and 5(b) can not create a good random numbers iid from uniform distribution (based on K-S test). But the combination generator of 5(c) can do it, because it has a longer cycle.
##
## One-sample Kolmogorov-Smirnov test
##
## data: res_5c
## D = 0.010053, p-value = 0.2643
## alternative hypothesis: two-sidedThe histograms of 5(a), 5(b) and 5(c) is shown below.
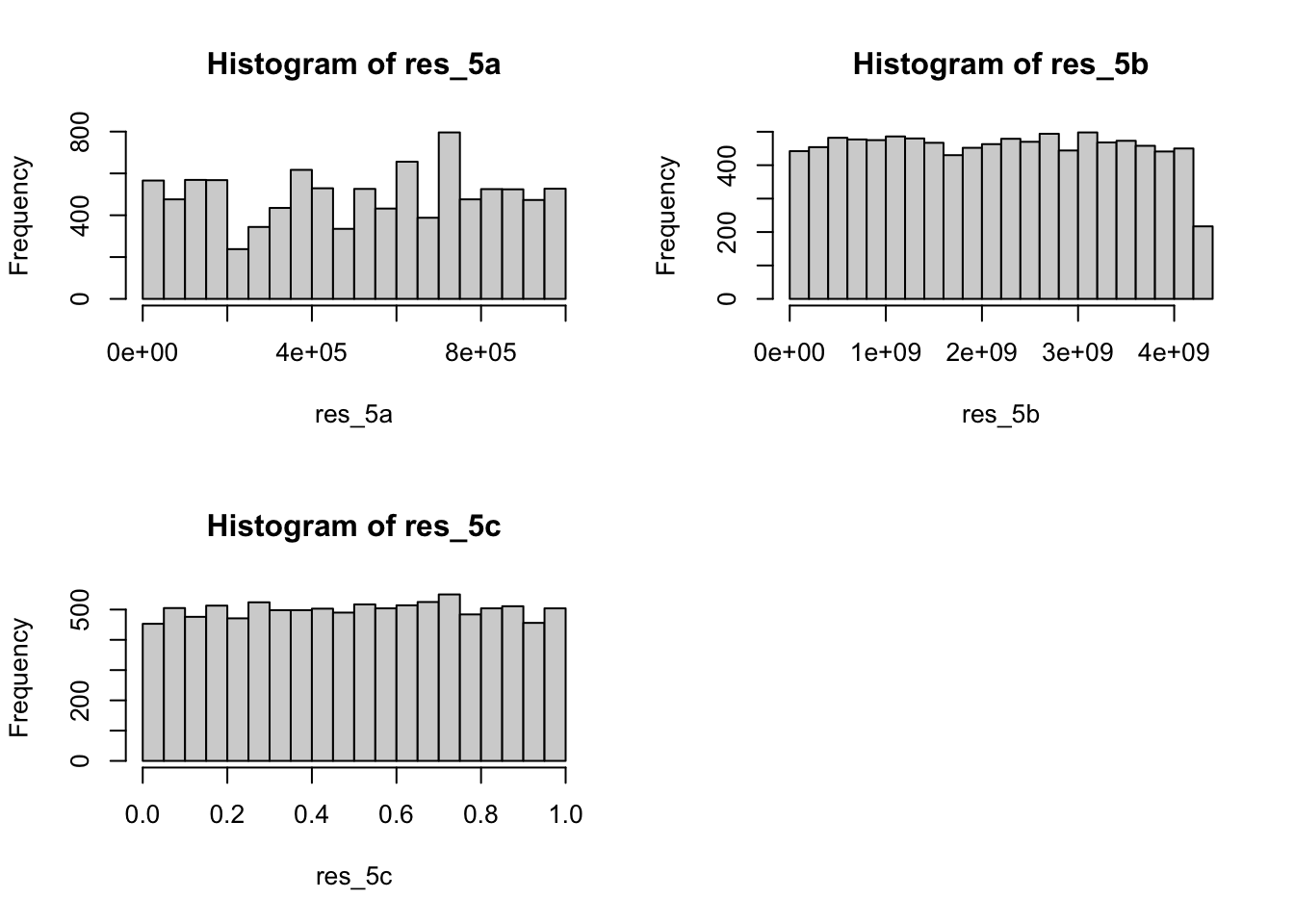
Ex. 6
(a) sample and ceiling(runif)
- We test a small value (=100) and a large value (=1000000).
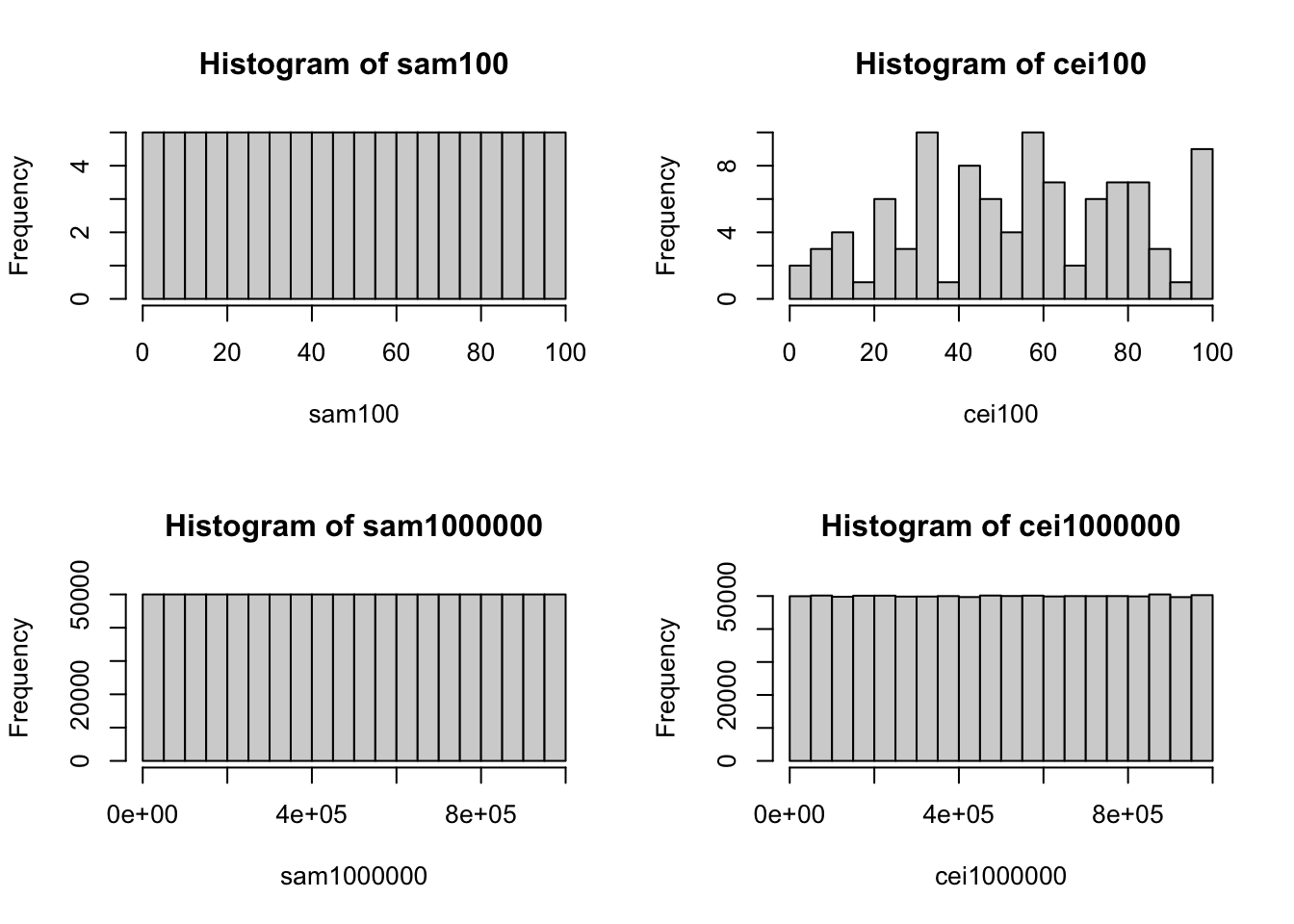
- Although these histograms above look very close to uniform distributions. But from the K-S tests, we can find any random numbers created by
sample()orceiling(runif())are not actually iid from uniform distributions.
##
## One-sample Kolmogorov-Smirnov test
##
## data: sam100
## D = 1, p-value < 2.2e-16
## alternative hypothesis: two-sided## Warning in ks.test(x = cei100, y = "punif"): ties should not be present for the
## Kolmogorov-Smirnov test##
## One-sample Kolmogorov-Smirnov test
##
## data: cei100
## D = 1, p-value < 2.2e-16
## alternative hypothesis: two-sided##
## One-sample Kolmogorov-Smirnov test
##
## data: sam1000000
## D = 1, p-value < 2.2e-16
## alternative hypothesis: two-sided## Warning in ks.test(x = cei1000000, y = "punif"): ties should not be present for
## the Kolmogorov-Smirnov test##
## One-sample Kolmogorov-Smirnov test
##
## data: cei1000000
## D = 1, p-value < 2.2e-16
## alternative hypothesis: two-sided(b) Hand calculators generator
- The comment of the
gen_6b()function. (1) Create a null vectorvec. (2) In the for loop, implement a generators based on uniform and \(\pi\)(pi+u)^5 %% 1. (3) Save the numbers in the vectorvecwith theith place. - Goodness-of-fit test. We generated 10000 random numbers from the
gen_6b()function. Check the K-S test D = 0.0090575, p = 0.3848, we can not reject the null hypothesis, i.e. the random numbers do follow the uniform distribution. - Compare to #5. In 5(c) we find only the combination generator can create random numbers followed the uniform distribution. Then we check independence by the runs test (from
snparpackage). The p-value of bothres_5corres_6bis at the 0.05 significance level, so we can conclude that the numbers form them are random (independent).
##
## One-sample Kolmogorov-Smirnov test
##
## data: res_6b
## D = 0.0090575, p-value = 0.3848
## alternative hypothesis: two-sided##
## Approximate runs rest
##
## data: res_5c
## Runs = 5019, p-value = 0.7188
## alternative hypothesis: two.sided##
## Approximate runs rest
##
## data: res_6b
## Runs = 4930, p-value = 0.1556
## alternative hypothesis: two.sided(c) \(\phi\) generator
- Replaced \(pi\) with \(\phi=\dfrac{1+\sqrt{5}}{2}\).
- Check the K-S test D = 0.0066451, p = 0.7692, we can not reject the null hypothesis, i.e. the random numbers do follow the uniform distribution. And from the runs test with p-value = 0.1676, we can find the numbers are random (independent).
##
## One-sample Kolmogorov-Smirnov test
##
## data: res_6c
## D = 0.0066451, p-value = 0.7692
## alternative hypothesis: two-sided##
## Approximate runs rest
##
## data: res_6c
## Runs = 4932, p-value = 0.1676
## alternative hypothesis: two.sided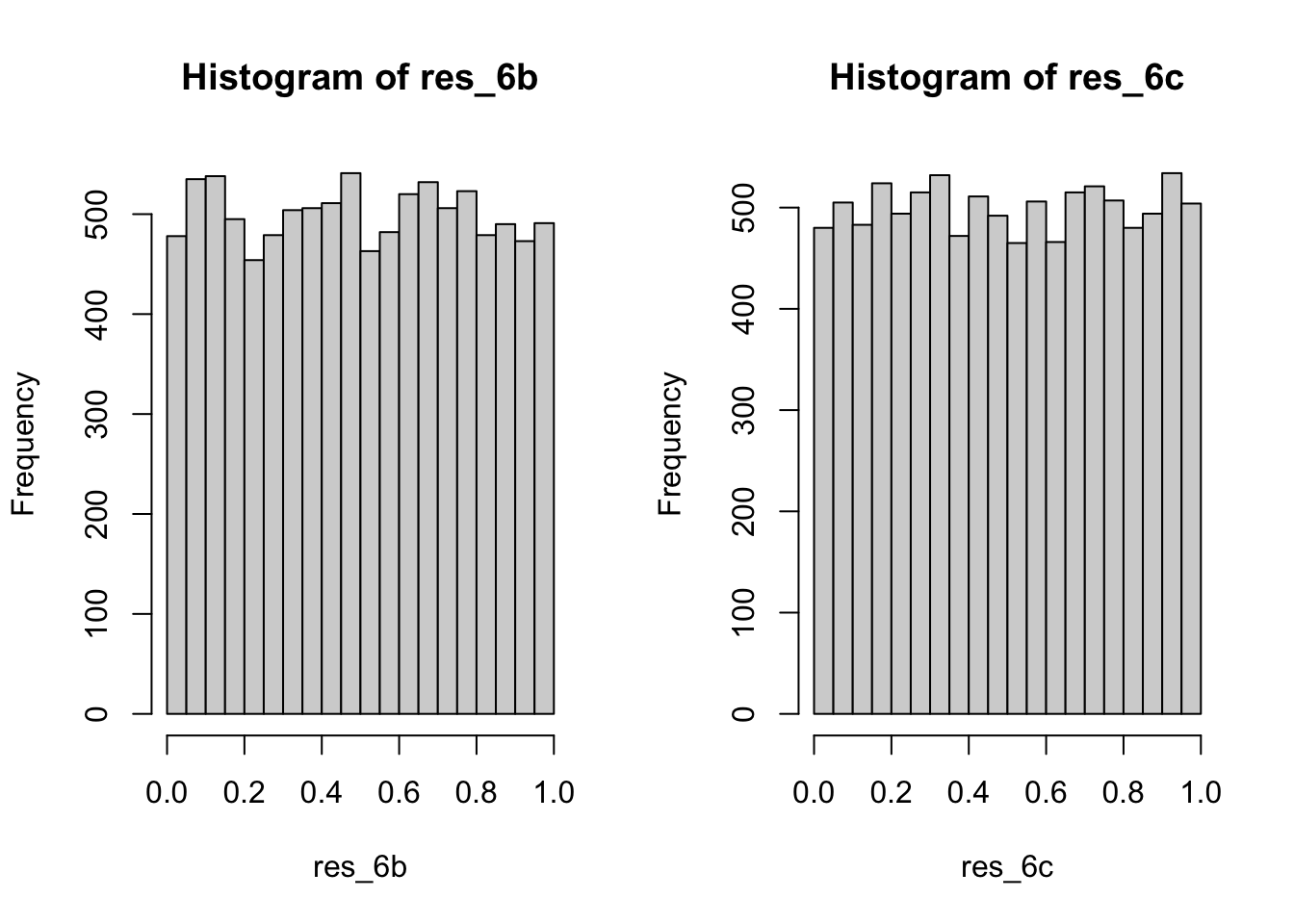
Appendix
Note. After R 4.0, some useful new features are built-in.
- The native pipeline syntax
|>replaced the tidy-style pipeline%>%. - The shorthand syntax for anonymous functions
\(x)is more compact thanfunction(x).
## 1(a) --------
dat1 <- scan('baseball-salaries-2016.txt', what = "character" , sep="\t")
dat1a <- matrix(dat1[-c(1:6)], ncol=6, byrow=T) |> data.frame()
names(dat1a) <- dat1[1:6]
# export data frame
write.csv(dat1a, 'dat1a.csv')
write.table(dat1a, 'dat1a.txt')
## 1(b) --------
dat1b <- rio::import('baseball-salaries-2016.txt')
par(mfrow=c(1,2))
boxplot((W/(W+L)) ~ League, data = dat1b, ylab = 'WPCT' )
boxplot(`Salary ($M)` ~ League, data = dat1b, ylab = 'Salary')
## 1(c) --------
sala <- dat1b$`Salary ($M)`
wpct <- dat1b$W/(dat1b$W+dat1b$L)
team <- dat1b$Team
cor.test(sala, wpct )
## 1(d) --------
x11()
plot(wpct~sala)
abline( lm(wpct~sala) , lwd=2)
identify(wpct~sala, labels = team)
## 2(a) --------
set.seed(1234)
x_2a = rnorm(20) |> round(2); print(x_2a, width = 65)
length(x_2a) = 15; print(x_2a, width = 65)
length(x_2a) = 18; print(x_2a, width = 65)
## 2(b) --------
set.seed(1234)
x_2a = rnorm(20) |> round(2)
x_2a[x_2a<0] <- NA
print(x_2a, width = 55)
paste0( 'mean: ',mean(x_2a[1:18], na.rm = T) |> round(2) )
paste0( 'median: ', median(x_2a[1:18], na.rm = T) |> round(2) )
paste0( 'var: ', var(x_2a[1:18], na.rm = T) |> round(2) )
## 2(c) --------
x_2a[is.na(x_2a)] <- 0; print(x_2a, width = 55)
## 3(a) 3(b) --------
eq1 <- \(x){ ifelse(x<=3, (x^2-2*x+3), NA) }
eq2 <- \(x){ ifelse(x>3,(x+2*exp(x-3)+1), NA) }
# draw the curves
curve(eq2, from=0, to=6, col='blue', lwd=2,
ylim = c(-5,50),
xlab="x", ylab="f(x)")
curve(eq1, from=0, to=6, col='red', lwd=2,
xlab="x", ylab="f(x)", add = T)
# add some lines
abline( h=2, lty=2 )
abline( b=(1+2*exp(2)), a=-8*exp(2)+1, lty=2 )
## 4(a) --------
cat(
labels = "Today’s date is: ",
paste( substr(date(),1,10), substr(date(),21,24)),
fill = TRUE
)
cat(
labels = "The time now is: ",
paste( substr(date(),12,19)),
fill = TRUE
)
## 5(a) --------
library('stringr')
# x= initial value, k=length, vec=random number vector
gen_5a <- \(x, k){
vec <- c()
for (i in 1:k){
x <- as.character(x^2) |>
stringr::str_pad( width=12, side='left', pad='0' ) |>
stringr::str_sub(4, 9) |>
as.numeric()
vec[i] <- x
}
return(vec)
}
set.seed(1234)
res_5a <- gen_5a( floor( runif(1)*(10^6) ), 10000 )
print( ks.test(res_5a, y="punif") )
## 5(b)--------
gen_5b <- \(x, k){
vec <- c()
for(i in 1:k){
x <- (69069*x) %% (2^32)
vec[i] <- x
}
return(vec)
}
set.seed(1234)
res_5b <- gen_5b( runif(1), k=10000 )
# ks test
print( ks.test(res_5b, y="punif") )
# chisq test
grp_5b <- \(x, k) {
int <- seq( min(x) , max(x) , (max(res_5b) - min(res_5b))/k )
vec <- c()
for (i in 1:k){
vec[i] <- sum( x<int[i+1] ) - sum( x<int[i] )
}
return(vec)
}
chi_5b <- grp_5b( res_5b, k=10 )
chisq.test( chi_5b, p= rep(1/10, 10) )
## 5(c) --------
gen_5c <- \(x, y, z){
vec <- c()
for(i in 1:10000) {
x <- (171*x) %% 30269
y <- (172*y) %% 30307
z <- (170*z) %% 30323
u <- (x/30269 + y/30307 + z/30323) %% 1
vec[i] <- u
}
return(vec)
}
res_5c <- gen_5c(x=1, y=2, z=3)
print( ks.test(res_5c, y="punif") )
# hist
par( mfrow=c(2,2) )
hist(res_5a, 20); hist(res_5b, 20); hist(res_5c, 20)
## 6(a) --------
set.seed(1234)
# 100
sam100 <- sample(100)
cei100 <- ceiling(runif(100)*100)
sam1000000 <- sample(1000000)
cei1000000 <- ceiling(runif(1000000)*1000000)
# graphical
par(mfrow=c(2,2))
hist(sam100, 20); hist(cei100, 20)
hist(sam1000000, 20); hist(cei1000000, 20)
# test
print(ks.test(x=sam100, y="punif"))
print(ks.test(x=cei100, y="punif"))
print(ks.test(x=sam1000000, y="punif"))
print(ks.test(x=cei1000000, y="punif"))
## 6(b) --------
set.seed(1234)
gen_6b <- \(k){
vec <- c()
u <- runif(1)
for(i in 1:k){
u <- (pi+u)^5 %% 1
vec[i] <- u
}
return(vec)
}
res_6b <- gen_6b(10000)
print( ks.test(res_6b, y="punif") )
snpar::runs.test( res_5c )
snpar::runs.test( res_6b )
# 6(c) --------
set.seed(1234)
gen_6c <- \(k){
vec <- c()
u <- runif(1)
phi <- (1+sqrt(5))/2
for(i in 1:k){
u <- (phi+u)^5 %% 1
vec[i] <- u
}
return(vec)
}
res_6c <- gen_6c(10000)
print( ks.test(res_6c, y="punif") )
snpar::runs.test( res_6c )
# hist
par(mfrow=c(1,2))
hist(res_6b, 20); hist(res_6c, 20)